Note
Go to the end to download the full example code.
Score-Then-Filter Graph Comparison Gallery#
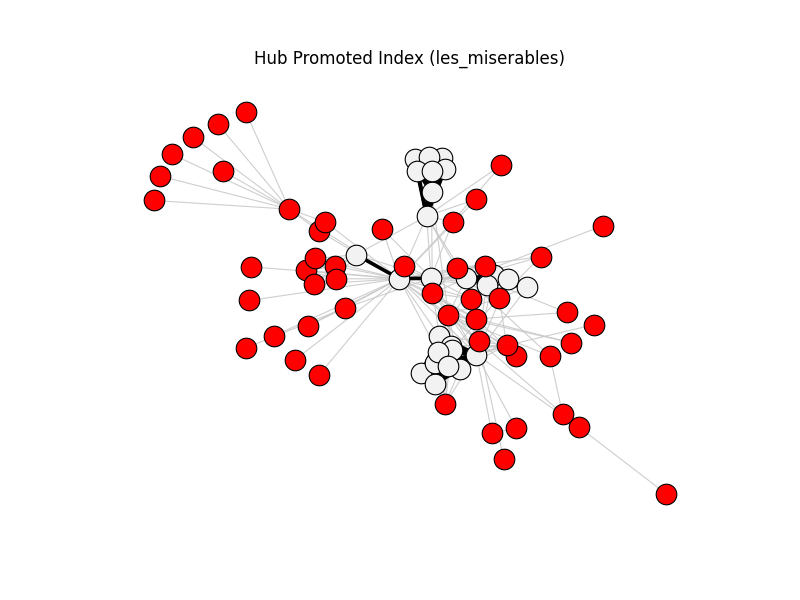
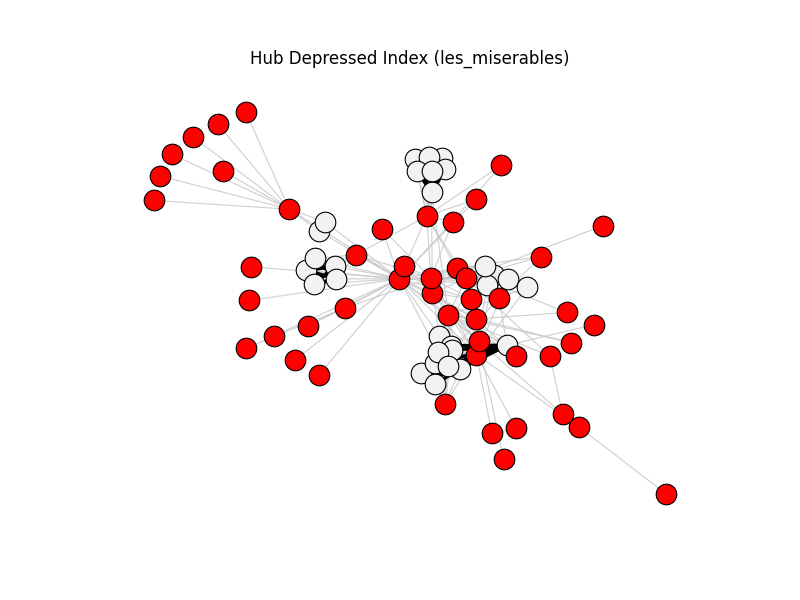
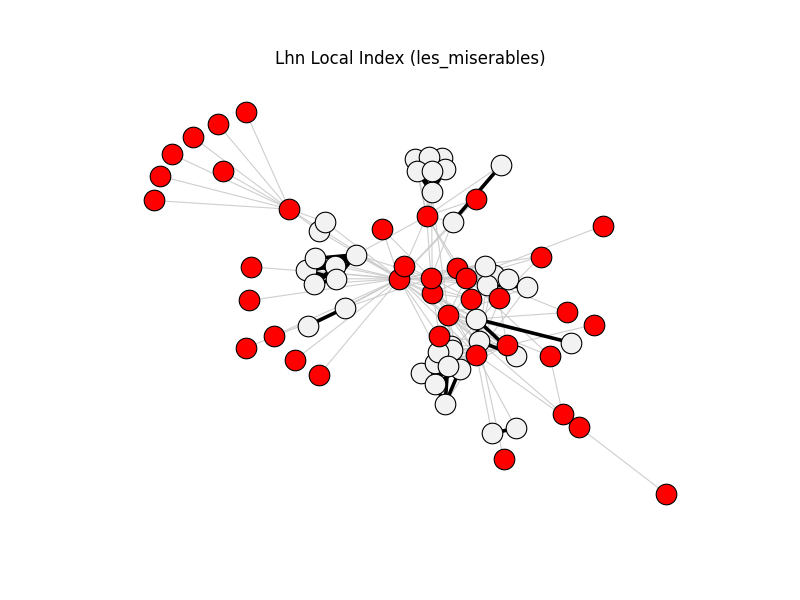
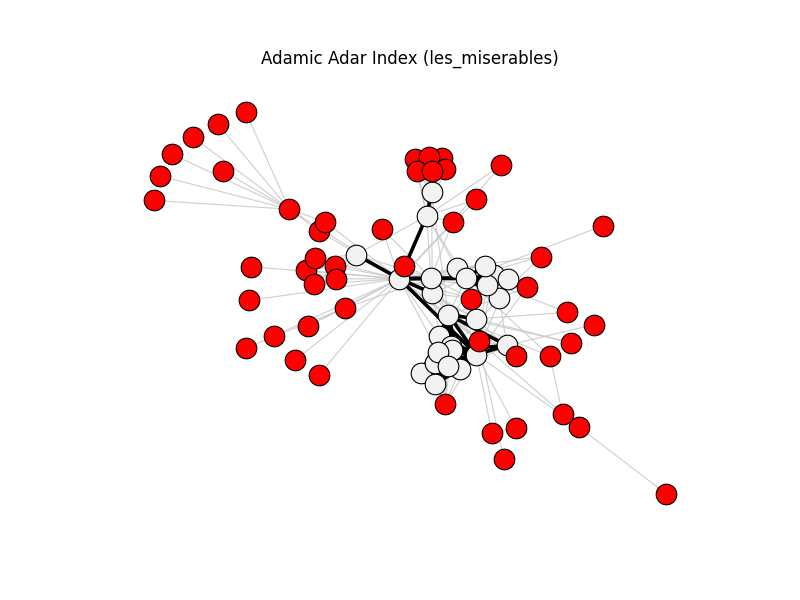
This gallery applies backbone methods to reference datasets, then visualizes filtered backbones against the original graphs.
For each method we:
Compute edge scores on the full graph.
Apply a filtering function to extract the backbone.
Warn if the filtered edge count is unchanged from the original graph.
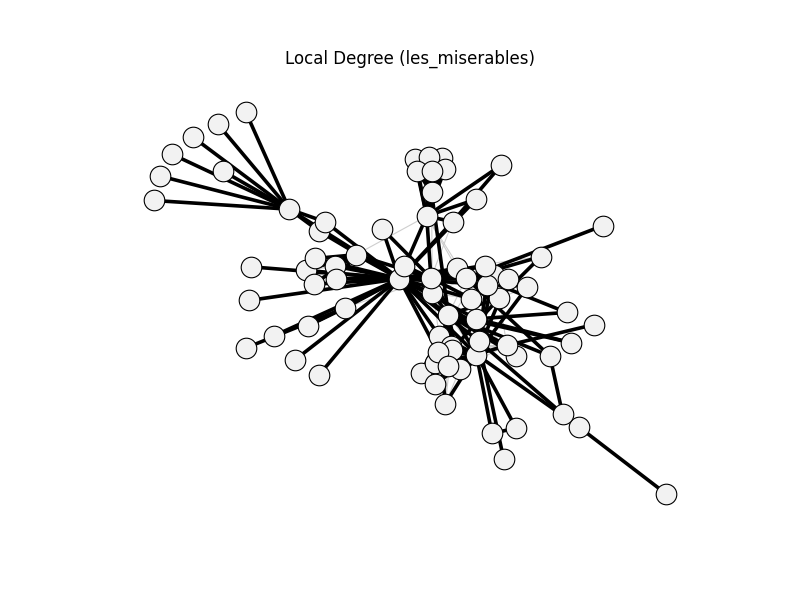
Non-bipartite methods use
nx.les_miserables_graph().Bipartite methods use
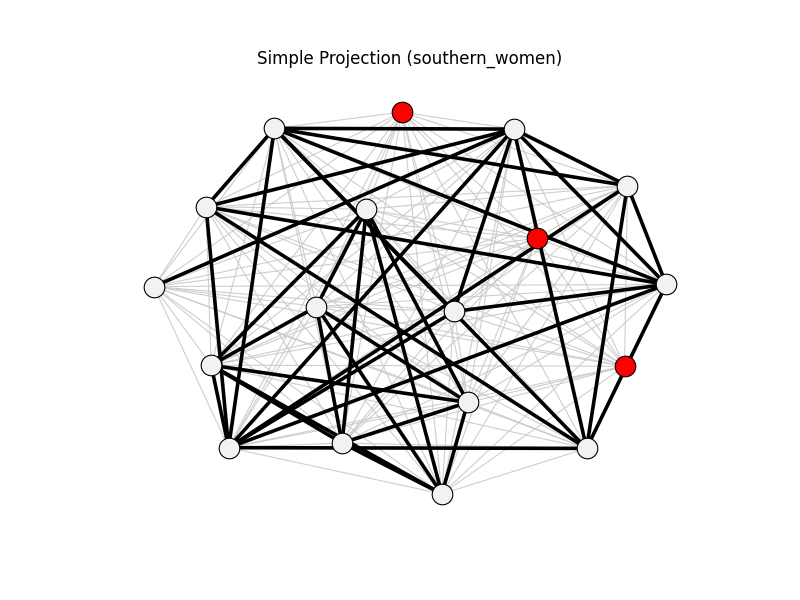
nx.davis_southern_women_graph()projected onto women nodes.
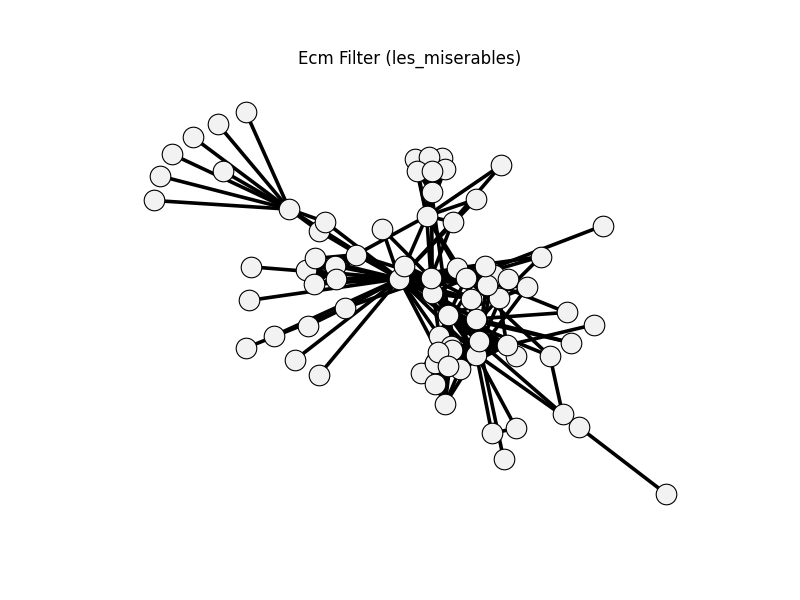
/home/runner/work/networkx_backbone/networkx_backbone/docs/examples/graph_comparison/plot_graph_comparison_gallery.py:596: UserWarning: ecm_filter (les_miserables) returned the same number of edges as the original graph. Re-test and validate threshold settings.
warnings.warn(
Method | Dataset | Original Edges | Backbone Edges | Removed Nodes
--- | --- | ---: | ---: | ---:
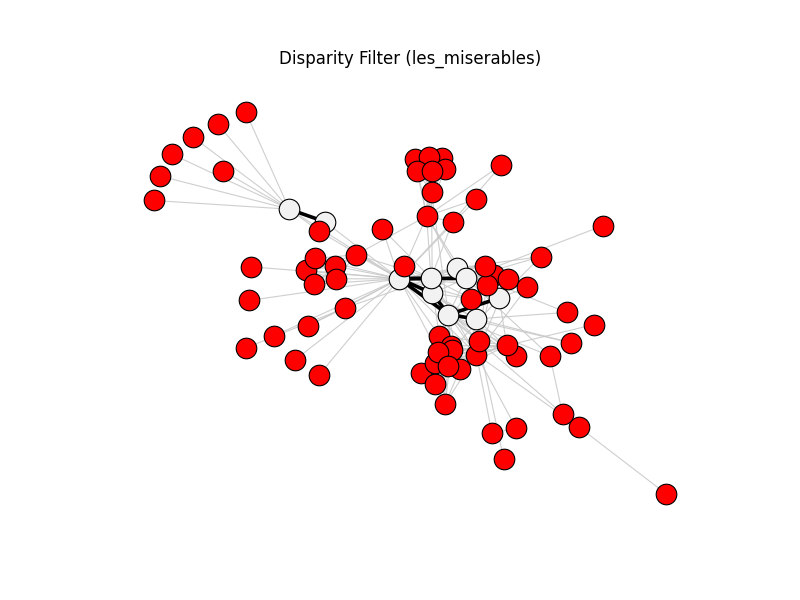
disparity_filter | les_miserables | 254 | 9 | 67
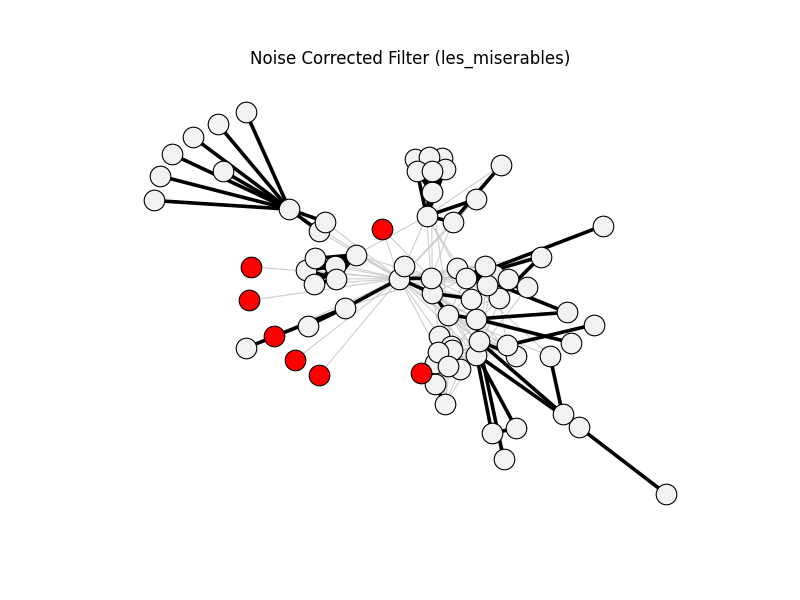
noise_corrected_filter | les_miserables | 254 | 98 | 7
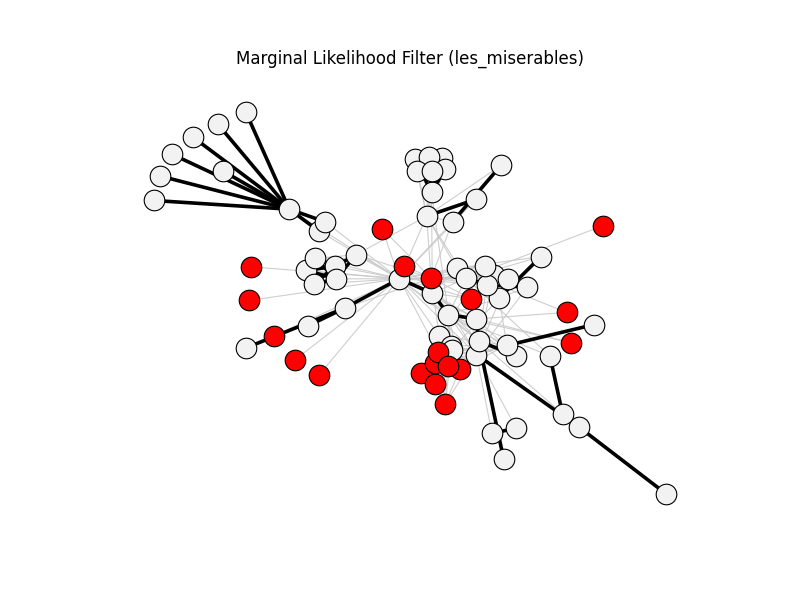
marginal_likelihood_filter | les_miserables | 254 | 70 | 19
ecm_filter | les_miserables | 254 | 254 | 0
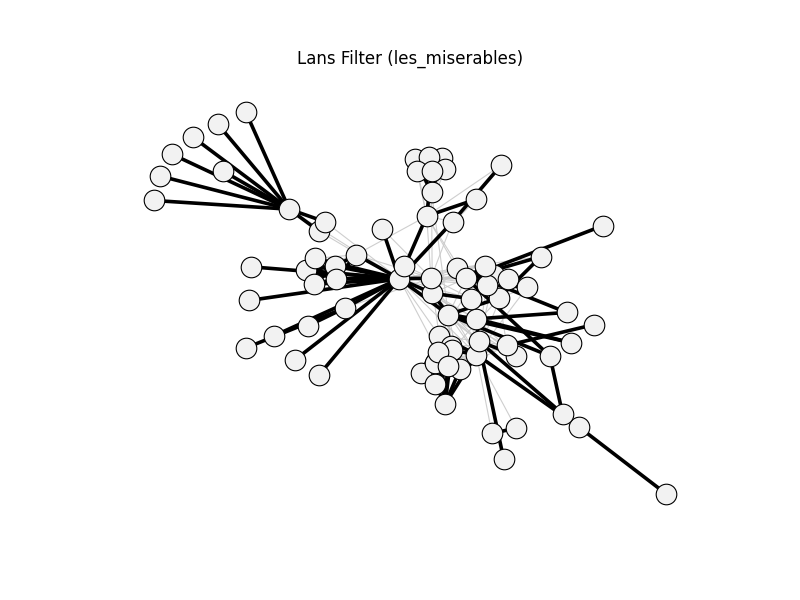
lans_filter | les_miserables | 254 | 109 | 0
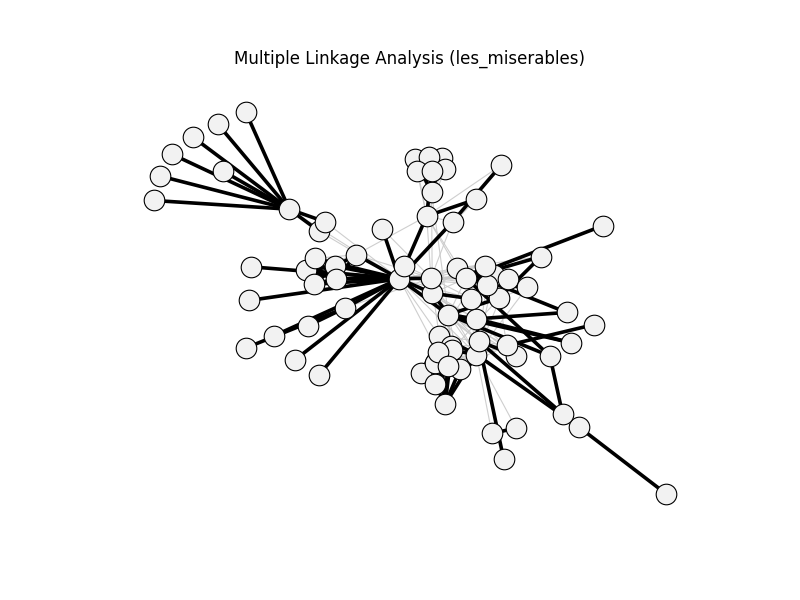
multiple_linkage_analysis | les_miserables | 254 | 109 | 0
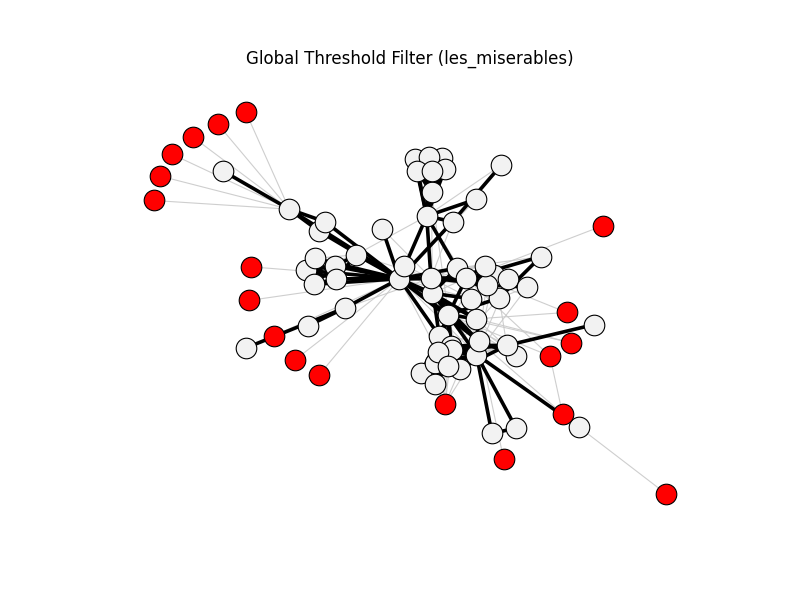
global_threshold_filter | les_miserables | 254 | 157 | 19
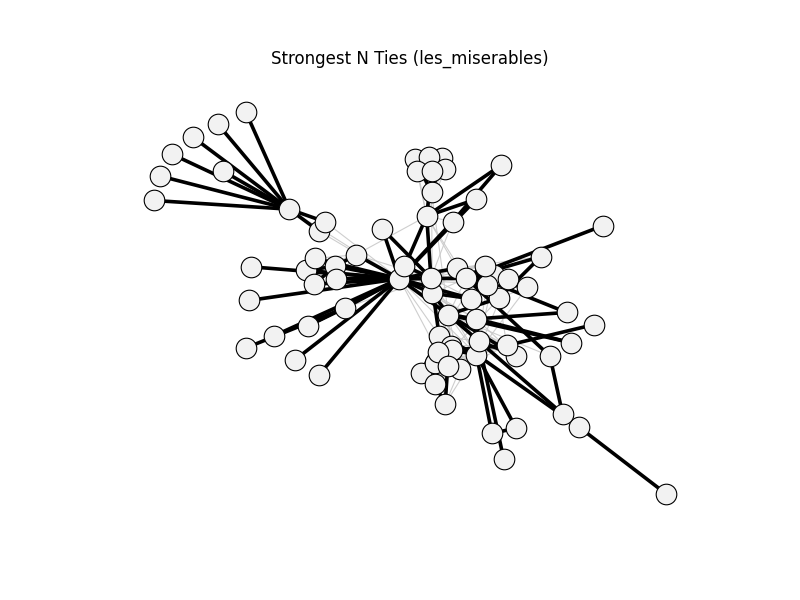
strongest_n_ties | les_miserables | 254 | 113 | 0
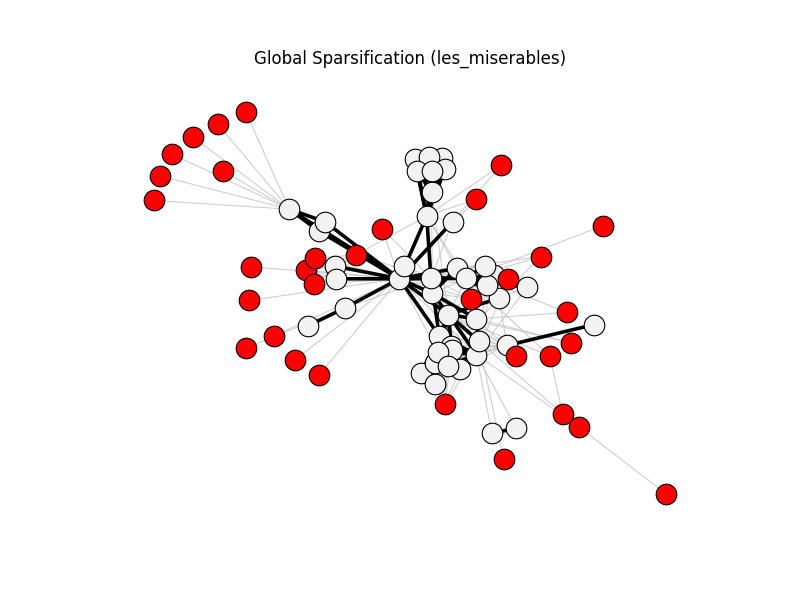
global_sparsification | les_miserables | 254 | 102 | 33
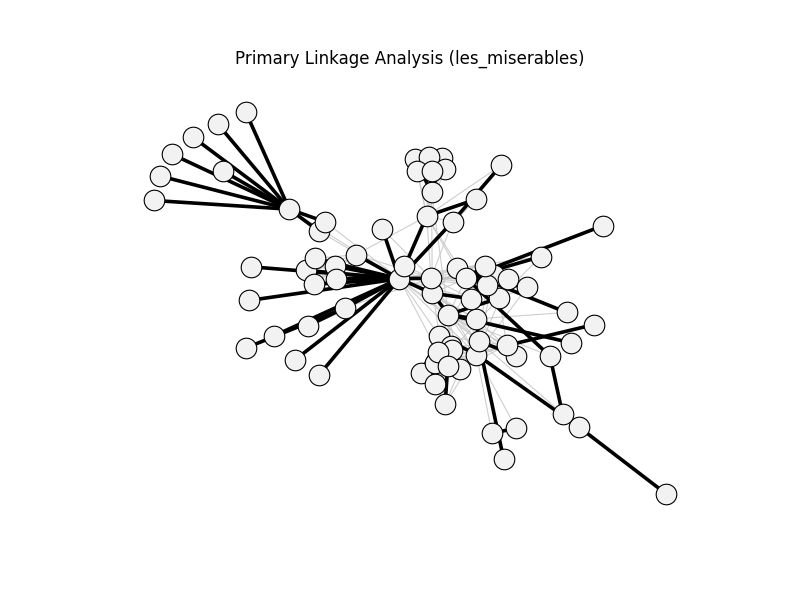
primary_linkage_analysis | les_miserables | 254 | 69 | 0
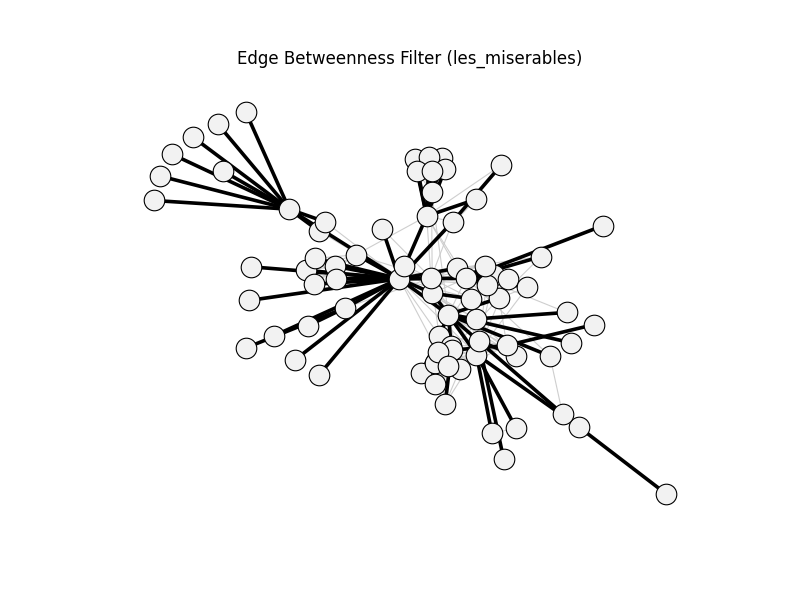
edge_betweenness_filter | les_miserables | 254 | 77 | 0
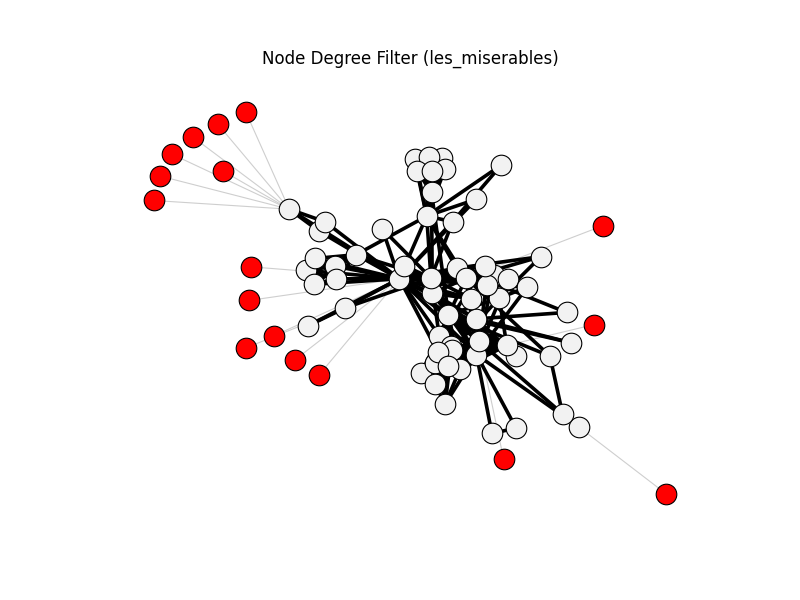
node_degree_filter | les_miserables | 254 | 237 | 17
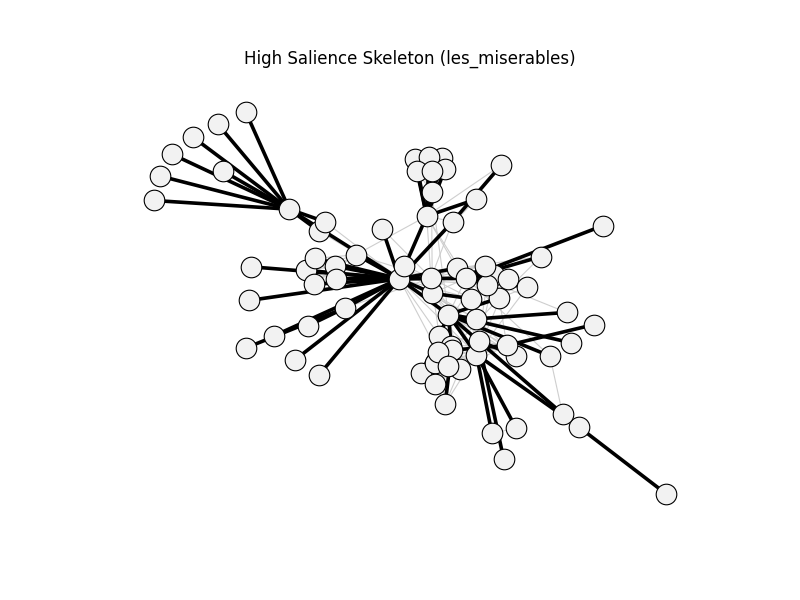
high_salience_skeleton | les_miserables | 254 | 76 | 0
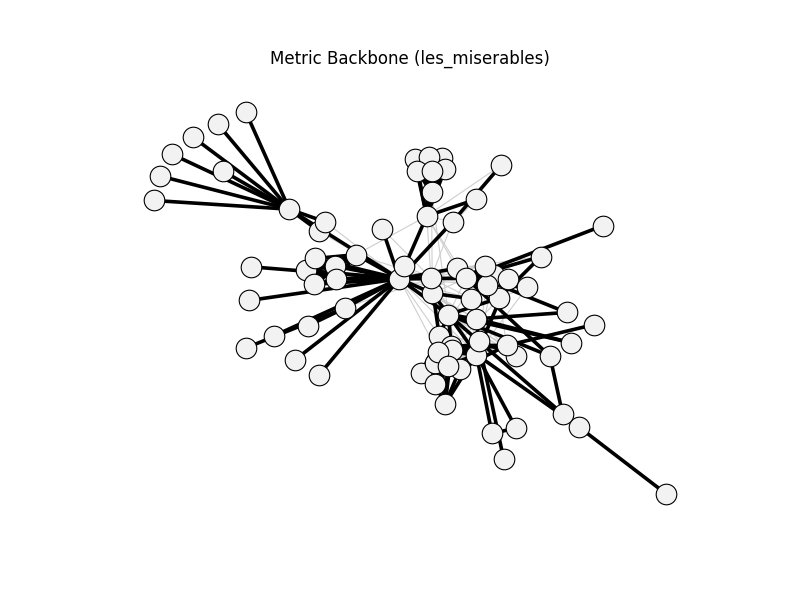
metric_backbone | les_miserables | 254 | 163 | 0
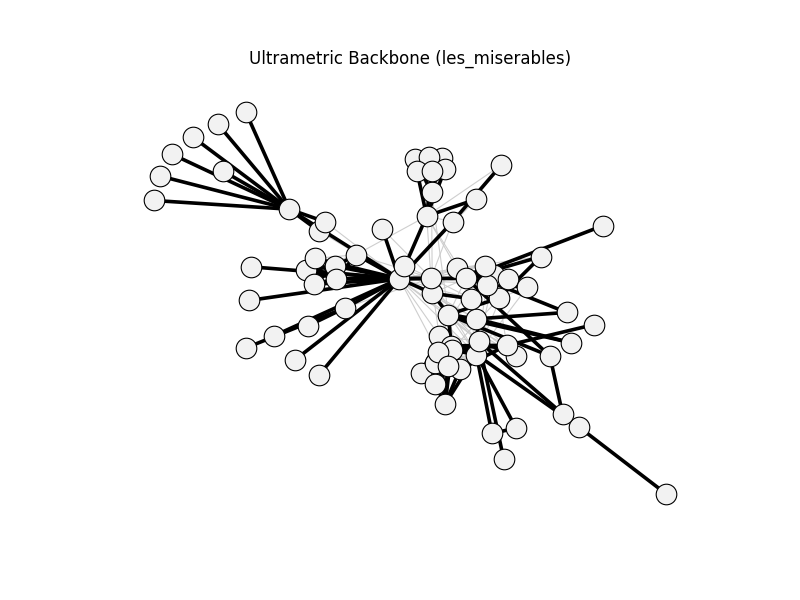
ultrametric_backbone | les_miserables | 254 | 118 | 0
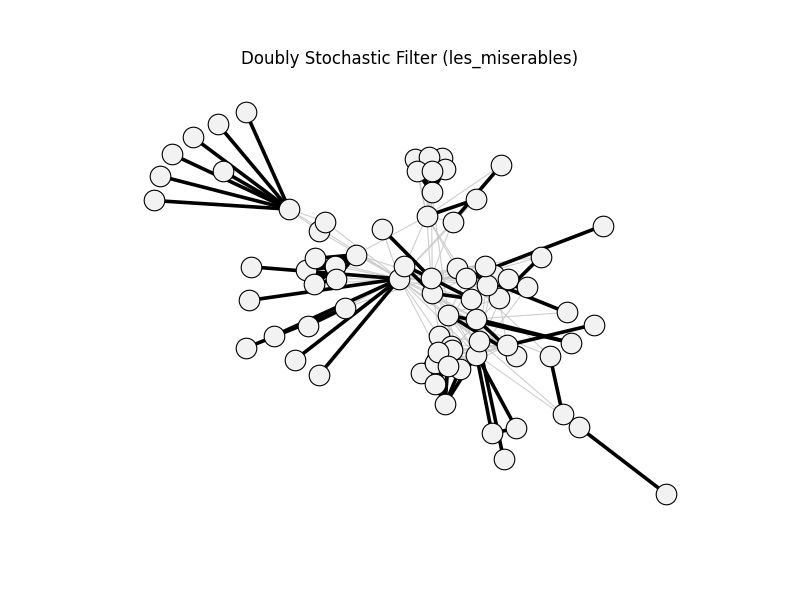
doubly_stochastic_filter | les_miserables | 254 | 109 | 0
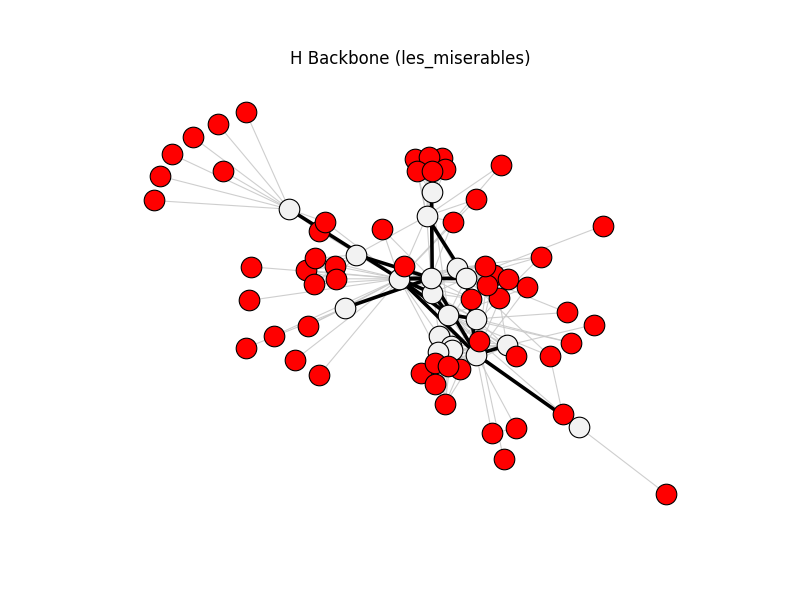
h_backbone | les_miserables | 254 | 22 | 58
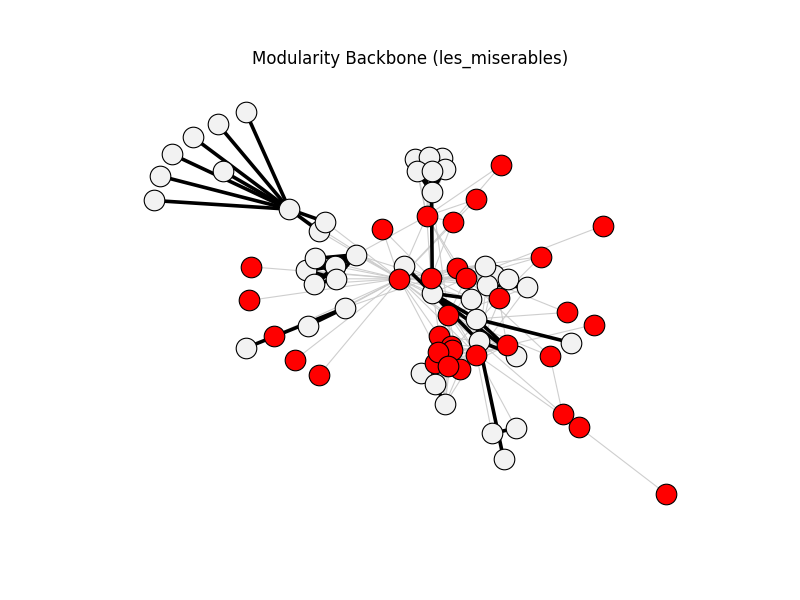
modularity_backbone | les_miserables | 254 | 72 | 33
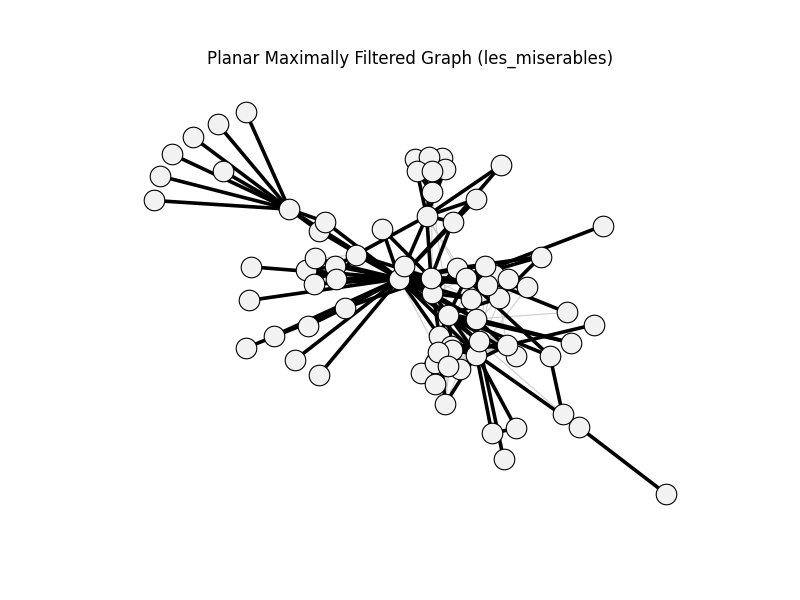
planar_maximally_filtered_graph | les_miserables | 254 | 162 | 0
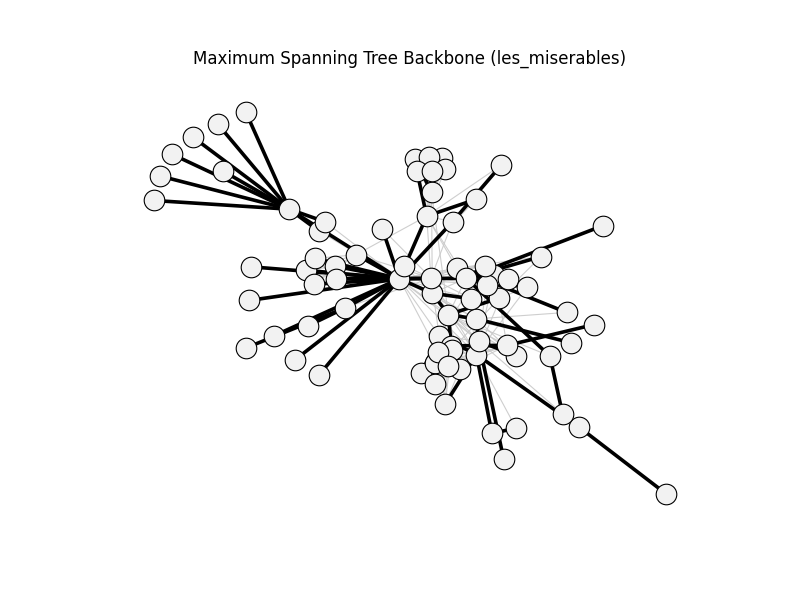
maximum_spanning_tree_backbone | les_miserables | 254 | 76 | 0
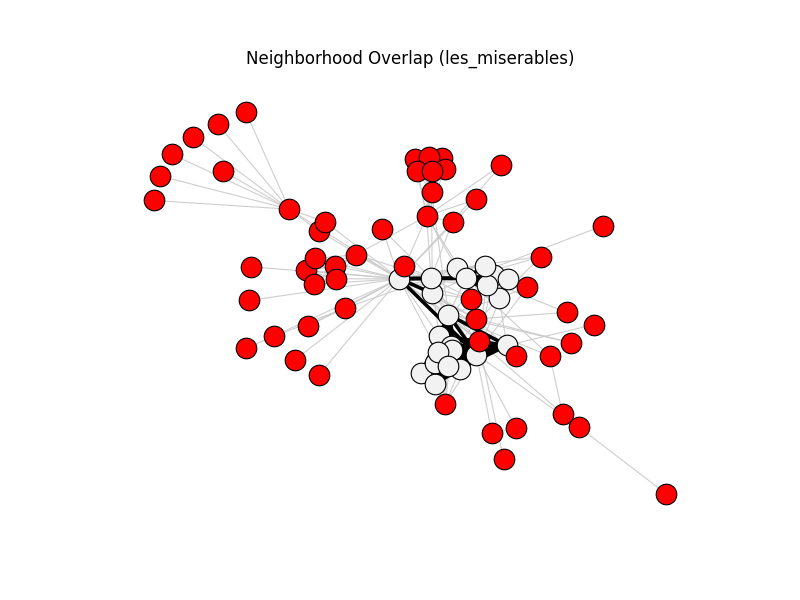
neighborhood_overlap | les_miserables | 254 | 76 | 55
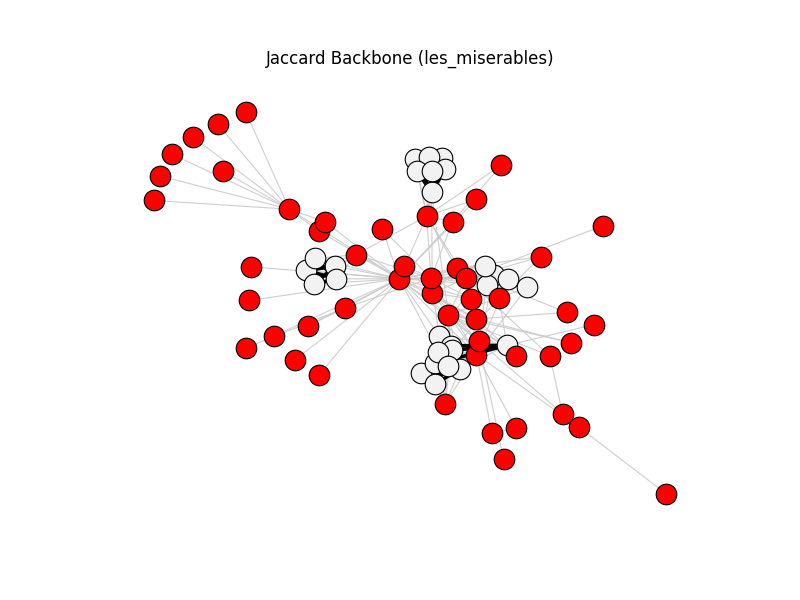
jaccard_backbone | les_miserables | 254 | 76 | 50
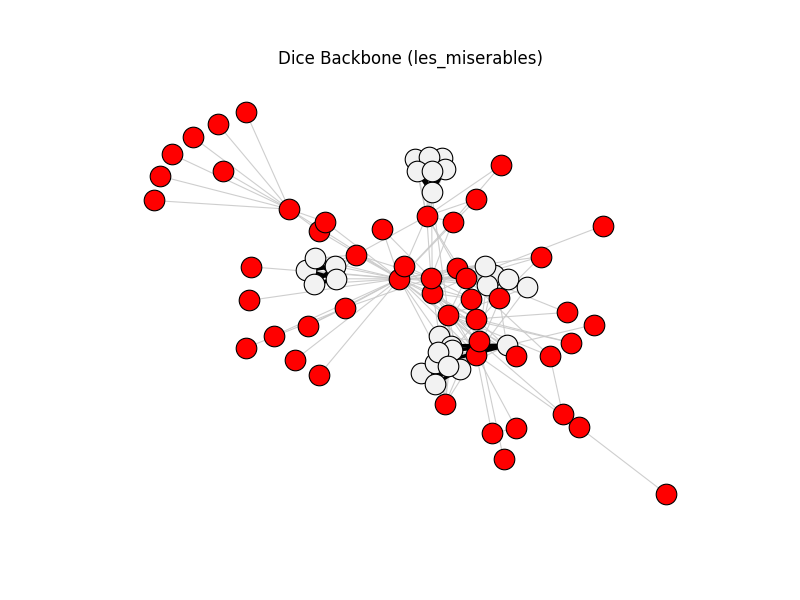
dice_backbone | les_miserables | 254 | 76 | 50
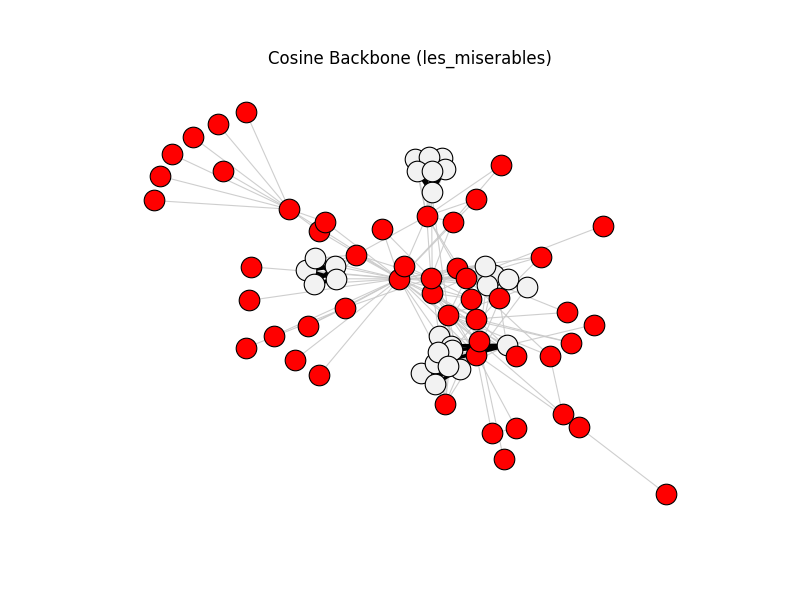
cosine_backbone | les_miserables | 254 | 76 | 50
hub_promoted_index | les_miserables | 254 | 76 | 51
hub_depressed_index | les_miserables | 254 | 76 | 48
lhn_local_index | les_miserables | 254 | 76 | 38
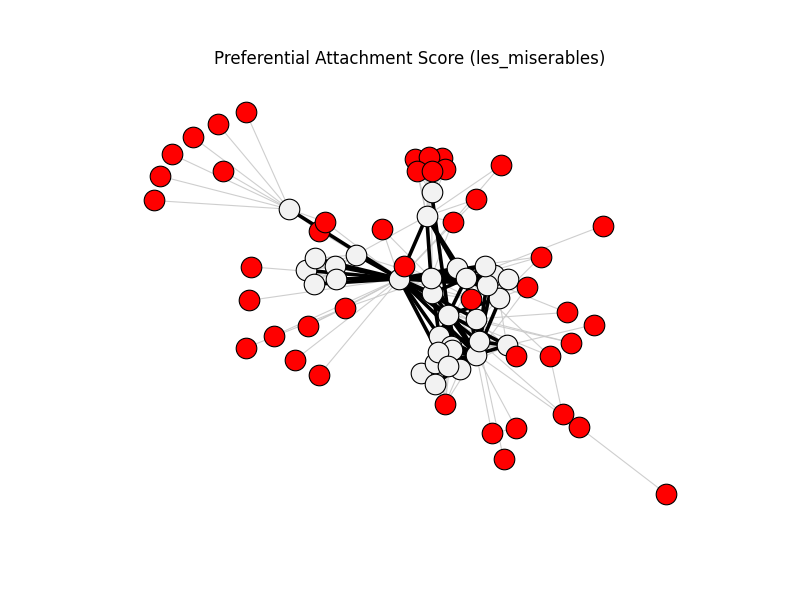
preferential_attachment_score | les_miserables | 254 | 76 | 44
adamic_adar_index | les_miserables | 254 | 76 | 51
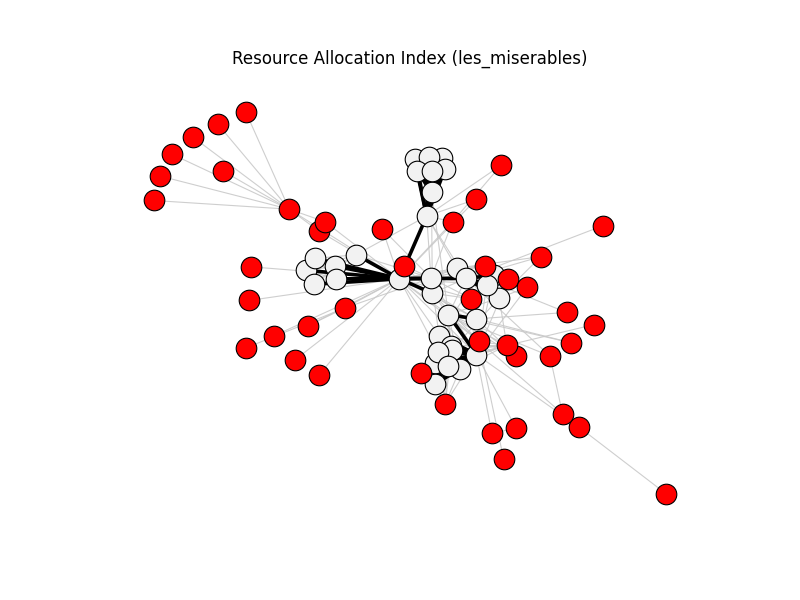
resource_allocation_index | les_miserables | 254 | 76 | 44
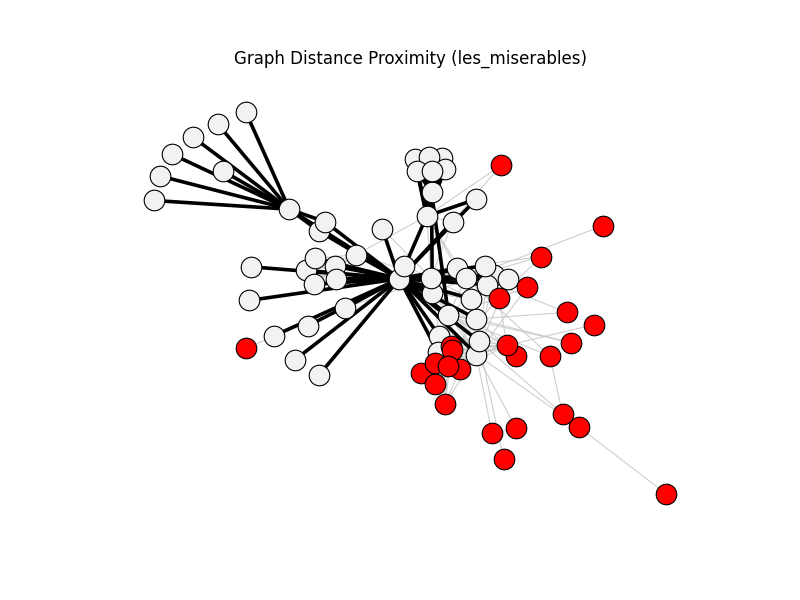
graph_distance_proximity | les_miserables | 254 | 76 | 26
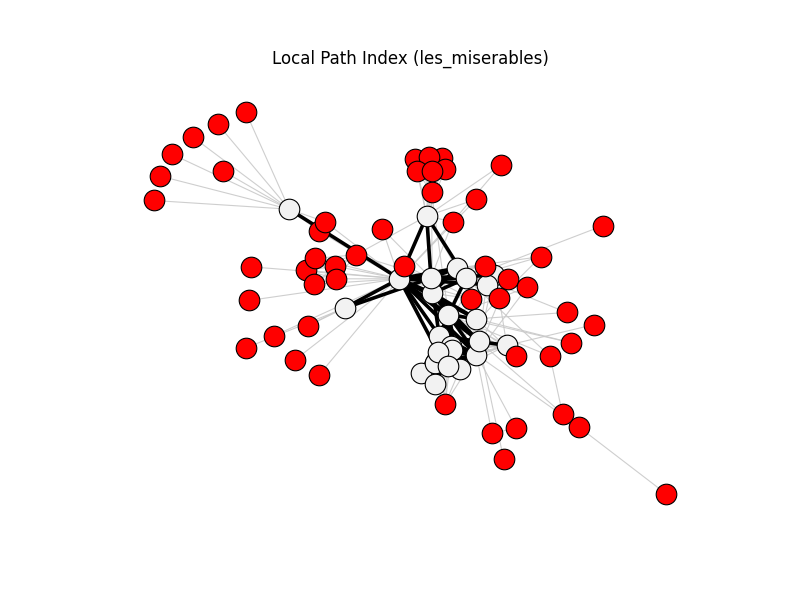
local_path_index | les_miserables | 254 | 76 | 53
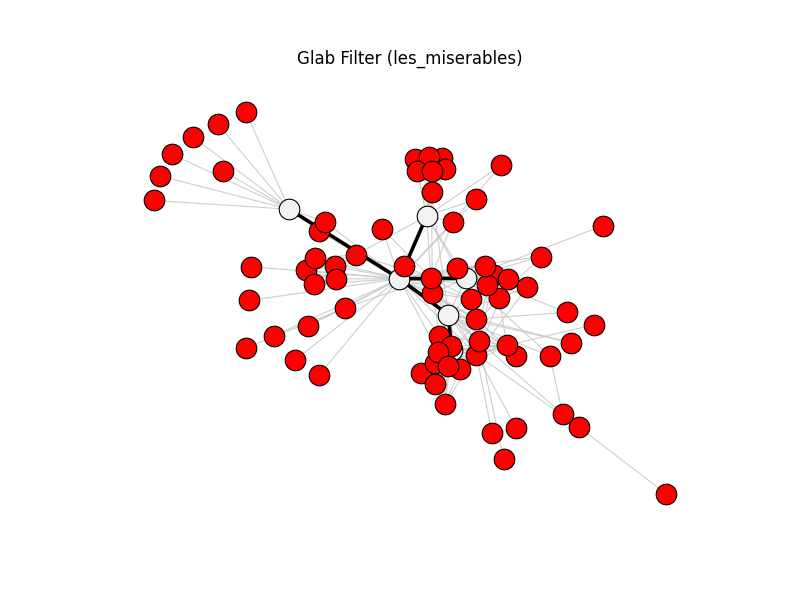
glab_filter | les_miserables | 254 | 5 | 71
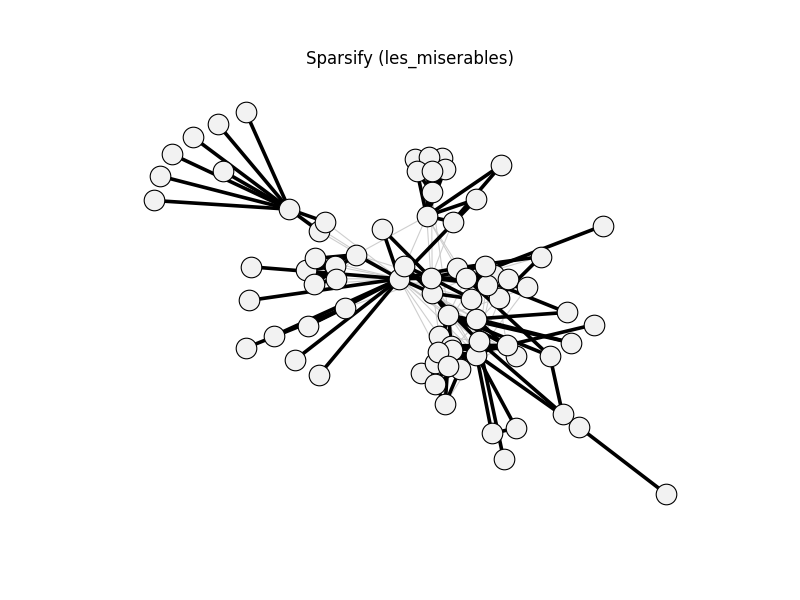
sparsify | les_miserables | 254 | 136 | 0
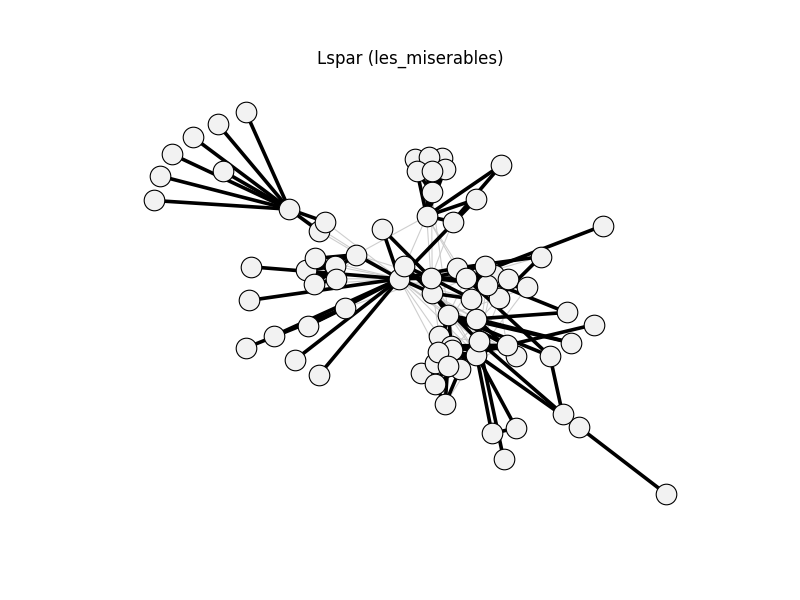
lspar | les_miserables | 254 | 136 | 0
local_degree | les_miserables | 254 | 135 | 0
simple_projection | southern_women | 153 | 41 | 3
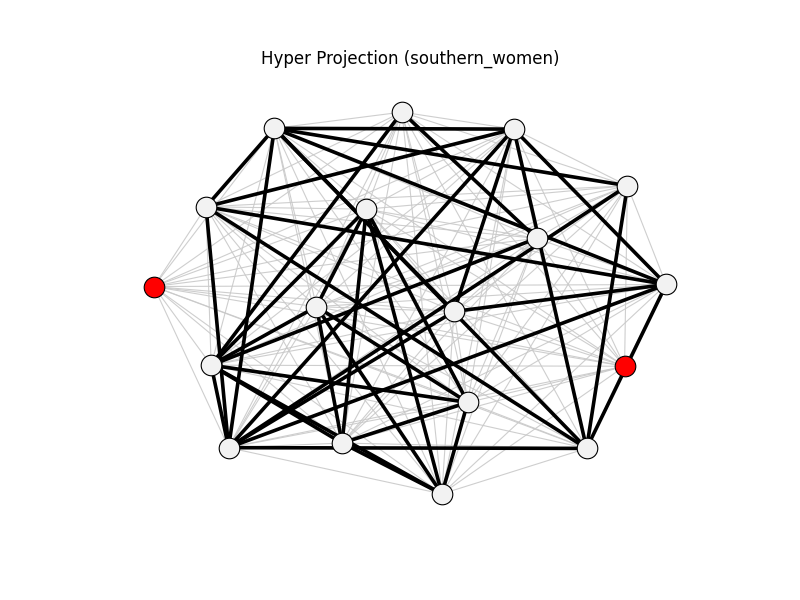
hyper_projection | southern_women | 153 | 41 | 2
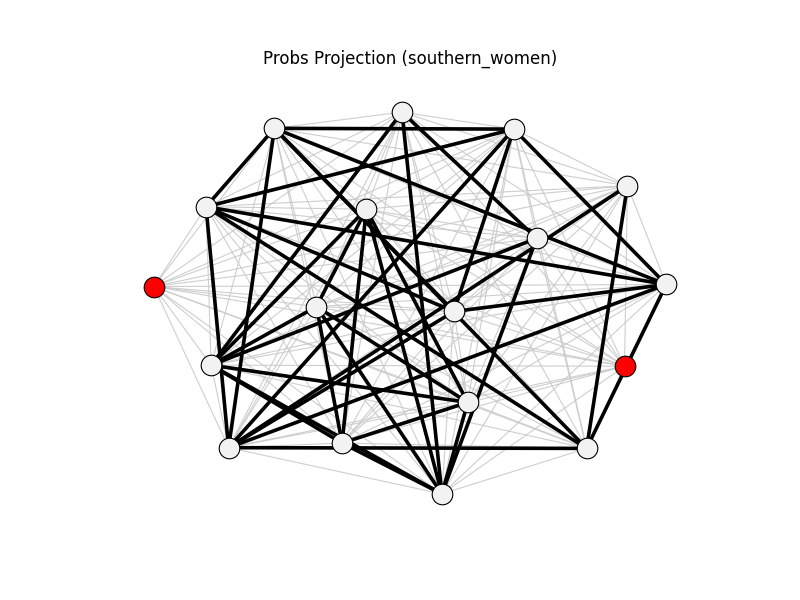
probs_projection | southern_women | 153 | 41 | 2
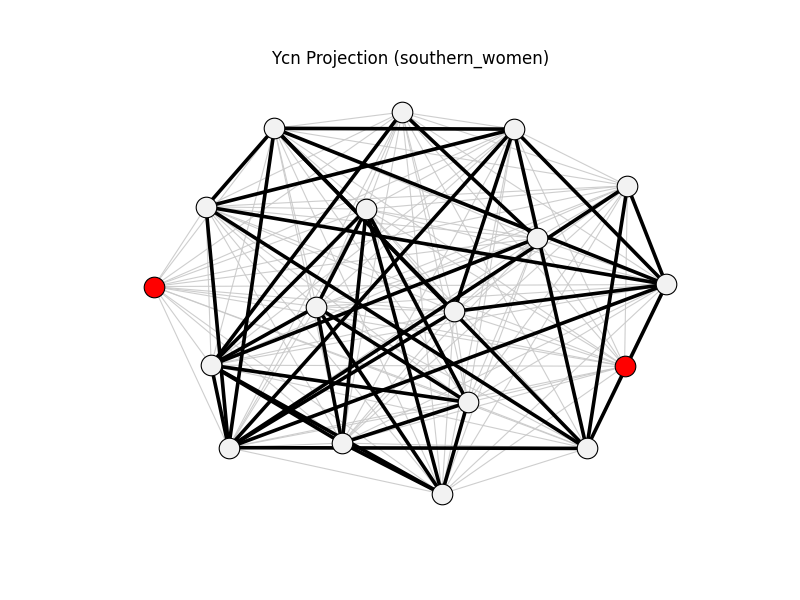
ycn_projection | southern_women | 153 | 41 | 2
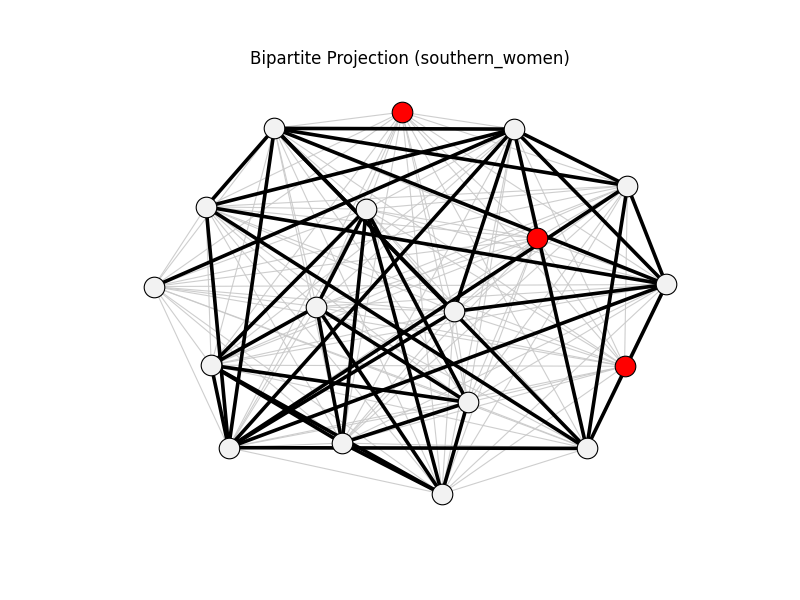
bipartite_projection | southern_women | 153 | 41 | 3
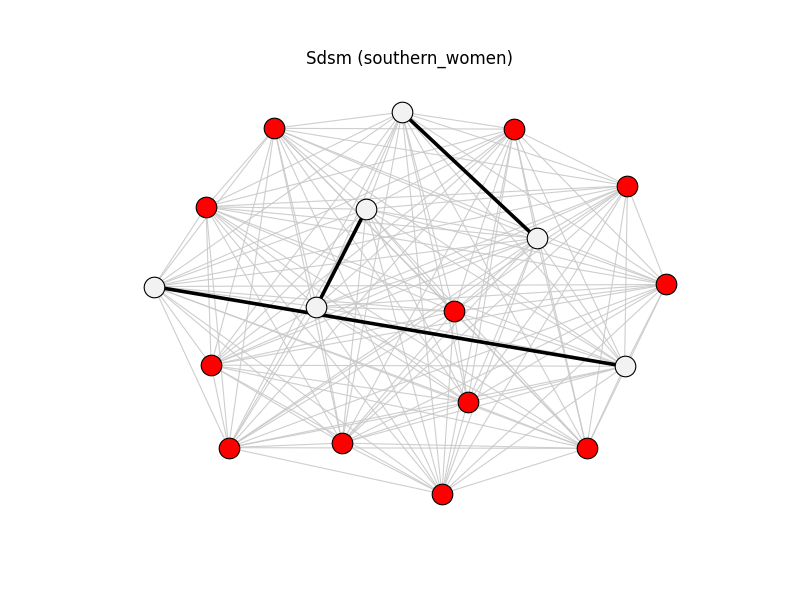
sdsm | southern_women | 153 | 3 | 12
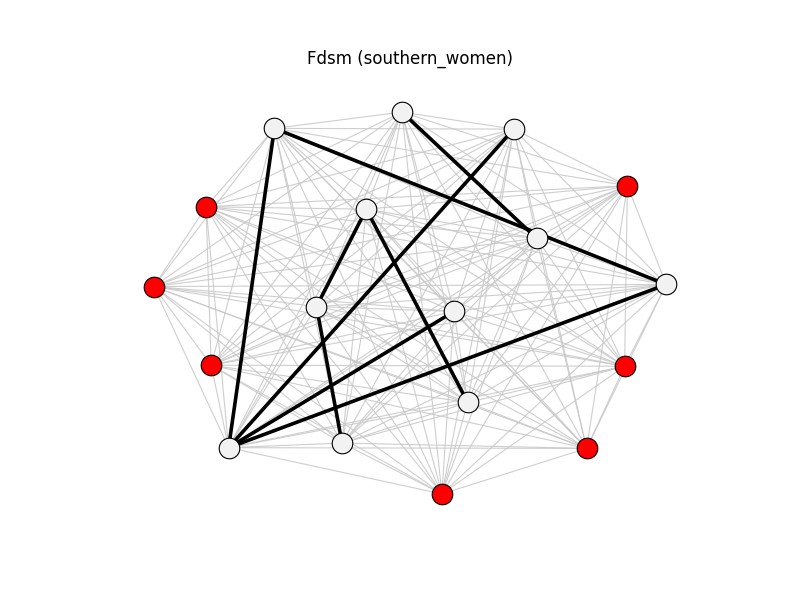
fdsm | southern_women | 153 | 9 | 7
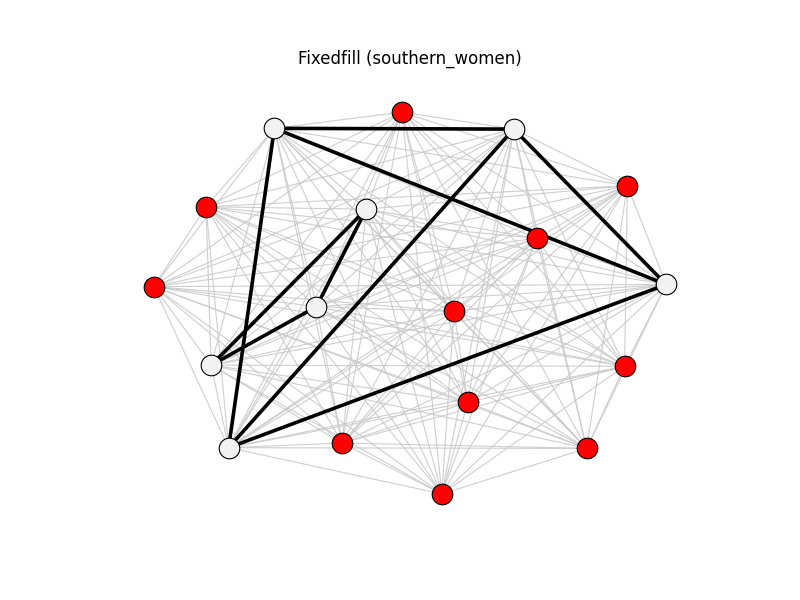
fixedfill | southern_women | 153 | 9 | 11
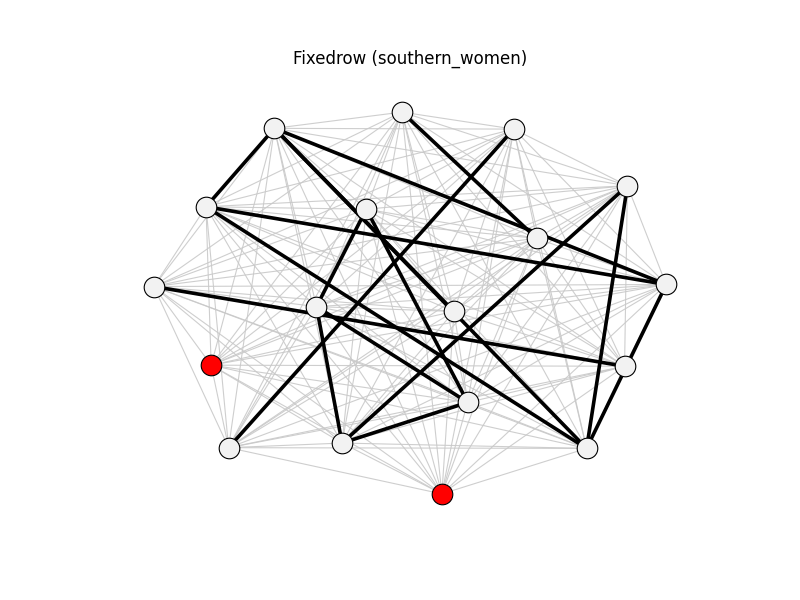
fixedrow | southern_women | 153 | 17 | 2
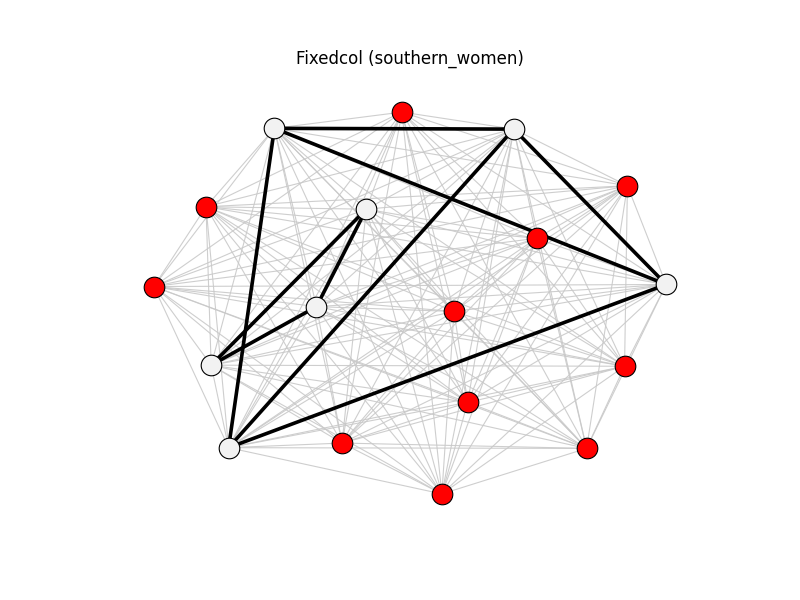
fixedcol | southern_women | 153 | 9 | 11
from __future__ import annotations
import warnings
from dataclasses import dataclass
from typing import Callable
import matplotlib.pyplot as plt
import networkx as nx
import networkx_backbone as nb
from networkx_backbone.visualization import compare_graphs
plt.rcParams["figure.max_open_warning"] = 0
@dataclass(frozen=True)
class MethodSpec:
name: str
dataset: str
module: str
build: Callable[[dict], nx.Graph]
def _to_unweighted(G: nx.Graph) -> nx.Graph:
H = nx.Graph()
H.add_nodes_from(G.nodes(data=True))
H.add_edges_from(G.edges())
return H
def _drop_isolates(G: nx.Graph) -> nx.Graph:
H = G.copy()
isolates = list(nx.isolates(H))
if isolates:
H.remove_nodes_from(isolates)
return H
def _to_undirected_simple(G: nx.Graph) -> nx.Graph:
if not G.is_directed():
return G
H = nx.Graph()
H.add_nodes_from(G.nodes(data=True))
for u, v, data in G.edges(data=True):
if not H.has_edge(u, v):
H.add_edge(u, v, **data)
return H
def _title(name: str) -> str:
return name.replace("_", " ").title()
def _method_specs() -> list[MethodSpec]:
return [
MethodSpec(
"disparity_filter",
"les_miserables",
"statistical",
lambda c: nb.threshold_filter(
nb.disparity_filter(c["les_weighted"]),
"disparity_pvalue",
0.05,
mode="below",
include_all_nodes=False,
),
),
MethodSpec(
"noise_corrected_filter",
"les_miserables",
"statistical",
lambda c: nb.threshold_filter(
nb.noise_corrected_filter(c["les_weighted"]),
"nc_score",
2.0,
mode="above",
include_all_nodes=False,
),
),
MethodSpec(
"marginal_likelihood_filter",
"les_miserables",
"statistical",
lambda c: nb.threshold_filter(
nb.marginal_likelihood_filter(c["les_weighted"]),
"ml_pvalue",
0.05,
mode="below",
include_all_nodes=False,
),
),
MethodSpec(
"ecm_filter",
"les_miserables",
"statistical",
lambda c: nb.threshold_filter(
nb.ecm_filter(c["les_weighted"]),
"ecm_pvalue",
0.05,
mode="below",
include_all_nodes=False,
),
),
MethodSpec(
"lans_filter",
"les_miserables",
"statistical",
lambda c: nb.threshold_filter(
nb.lans_filter(c["les_weighted"]),
"lans_pvalue",
0.05,
mode="below",
include_all_nodes=False,
),
),
MethodSpec(
"multiple_linkage_analysis",
"les_miserables",
"statistical",
lambda c: nb.boolean_filter(
nb.multiple_linkage_analysis(c["les_weighted"], alpha=0.05),
"mla_keep",
),
),
MethodSpec(
"global_threshold_filter",
"les_miserables",
"structural",
lambda c: nb.boolean_filter(
nb.global_threshold_filter(c["les_weighted"], threshold=2.0),
"global_threshold_keep",
),
),
MethodSpec(
"strongest_n_ties",
"les_miserables",
"structural",
lambda c: nb.boolean_filter(
nb.strongest_n_ties(c["les_weighted"], n=2),
"strongest_n_ties_keep",
),
),
MethodSpec(
"global_sparsification",
"les_miserables",
"structural",
lambda c: nb.boolean_filter(
nb.global_sparsification(c["les_weighted"], s=0.4),
"global_sparsification_keep",
),
),
MethodSpec(
"primary_linkage_analysis",
"les_miserables",
"structural",
lambda c: nb.boolean_filter(
nb.primary_linkage_analysis(c["les_weighted"]),
"primary_linkage_keep",
),
),
MethodSpec(
"edge_betweenness_filter",
"les_miserables",
"structural",
lambda c: nb.boolean_filter(
nb.edge_betweenness_filter(c["les_weighted"], s=0.3),
"edge_betweenness_keep",
),
),
MethodSpec(
"node_degree_filter",
"les_miserables",
"structural",
lambda c: nb.boolean_filter(
nb.node_degree_filter(c["les_weighted"], min_degree=2),
"node_degree_keep",
),
),
MethodSpec(
"high_salience_skeleton",
"les_miserables",
"structural",
lambda c: nb.threshold_filter(
nb.high_salience_skeleton(c["les_weighted"]),
"salience",
0.5,
mode="above",
include_all_nodes=False,
),
),
MethodSpec(
"metric_backbone",
"les_miserables",
"structural",
lambda c: nb.boolean_filter(
nb.metric_backbone(c["les_weighted"]),
"metric_keep",
),
),
MethodSpec(
"ultrametric_backbone",
"les_miserables",
"structural",
lambda c: nb.boolean_filter(
nb.ultrametric_backbone(c["les_weighted"]),
"ultrametric_keep",
),
),
MethodSpec(
"doubly_stochastic_filter",
"les_miserables",
"structural",
lambda c: nb.threshold_filter(
nb.doubly_stochastic_filter(c["les_weighted"]),
"ds_weight",
0.1,
mode="above",
include_all_nodes=False,
),
),
MethodSpec(
"h_backbone",
"les_miserables",
"structural",
lambda c: nb.boolean_filter(
nb.h_backbone(c["les_weighted"]),
"h_backbone_keep",
),
),
MethodSpec(
"modularity_backbone",
"les_miserables",
"structural",
lambda c: nb.boolean_filter(
nb.modularity_backbone(c["les_weighted"]),
"modularity_keep",
),
),
MethodSpec(
"planar_maximally_filtered_graph",
"les_miserables",
"structural",
lambda c: nb.boolean_filter(
nb.planar_maximally_filtered_graph(c["les_weighted"]),
"pmfg_keep",
),
),
MethodSpec(
"maximum_spanning_tree_backbone",
"les_miserables",
"structural",
lambda c: nb.boolean_filter(
nb.maximum_spanning_tree_backbone(c["les_weighted"]),
"mst_keep",
),
),
MethodSpec(
"neighborhood_overlap",
"les_miserables",
"proximity",
lambda c: nb.fraction_filter(
nb.neighborhood_overlap(c["les_weighted"]),
"overlap",
0.3,
ascending=False,
),
),
MethodSpec(
"jaccard_backbone",
"les_miserables",
"proximity",
lambda c: nb.fraction_filter(
nb.jaccard_backbone(c["les_weighted"]),
"jaccard",
0.3,
ascending=False,
),
),
MethodSpec(
"dice_backbone",
"les_miserables",
"proximity",
lambda c: nb.fraction_filter(
nb.dice_backbone(c["les_weighted"]),
"dice",
0.3,
ascending=False,
),
),
MethodSpec(
"cosine_backbone",
"les_miserables",
"proximity",
lambda c: nb.fraction_filter(
nb.cosine_backbone(c["les_weighted"]),
"cosine",
0.3,
ascending=False,
),
),
MethodSpec(
"hub_promoted_index",
"les_miserables",
"proximity",
lambda c: nb.fraction_filter(
nb.hub_promoted_index(c["les_weighted"]),
"hpi",
0.3,
ascending=False,
),
),
MethodSpec(
"hub_depressed_index",
"les_miserables",
"proximity",
lambda c: nb.fraction_filter(
nb.hub_depressed_index(c["les_weighted"]),
"hdi",
0.3,
ascending=False,
),
),
MethodSpec(
"lhn_local_index",
"les_miserables",
"proximity",
lambda c: nb.fraction_filter(
nb.lhn_local_index(c["les_weighted"]),
"lhn_local",
0.3,
ascending=False,
),
),
MethodSpec(
"preferential_attachment_score",
"les_miserables",
"proximity",
lambda c: nb.fraction_filter(
nb.preferential_attachment_score(c["les_weighted"]),
"pa",
0.3,
ascending=False,
),
),
MethodSpec(
"adamic_adar_index",
"les_miserables",
"proximity",
lambda c: nb.fraction_filter(
nb.adamic_adar_index(c["les_weighted"]),
"adamic_adar",
0.3,
ascending=False,
),
),
MethodSpec(
"resource_allocation_index",
"les_miserables",
"proximity",
lambda c: nb.fraction_filter(
nb.resource_allocation_index(c["les_weighted"]),
"resource_allocation",
0.3,
ascending=False,
),
),
MethodSpec(
"graph_distance_proximity",
"les_miserables",
"proximity",
lambda c: nb.fraction_filter(
nb.graph_distance_proximity(c["les_weighted"], all_pairs=False),
"dist",
0.3,
ascending=False,
),
),
MethodSpec(
"local_path_index",
"les_miserables",
"proximity",
lambda c: nb.fraction_filter(
nb.local_path_index(c["les_weighted"]),
"lp",
0.3,
ascending=False,
),
),
MethodSpec(
"glab_filter",
"les_miserables",
"hybrid",
lambda c: nb.threshold_filter(
nb.glab_filter(c["les_weighted"]),
"glab_pvalue",
0.05,
mode="below",
include_all_nodes=False,
),
),
MethodSpec(
"sparsify",
"les_miserables",
"unweighted",
lambda c: nb.boolean_filter(
nb.sparsify(c["les_unweighted"], s=0.5),
"sparsify_keep",
),
),
MethodSpec(
"lspar",
"les_miserables",
"unweighted",
lambda c: nb.boolean_filter(
nb.lspar(c["les_unweighted"], s=0.5),
"sparsify_keep",
),
),
MethodSpec(
"local_degree",
"les_miserables",
"unweighted",
lambda c: nb.boolean_filter(
nb.local_degree(c["les_unweighted"], s=0.3),
"sparsify_keep",
),
),
MethodSpec(
"simple_projection",
"southern_women",
"bipartite",
lambda c: nb.fraction_filter(
nb.simple_projection(c["davis_bipartite"], c["women_nodes"]),
"weight",
0.3,
ascending=False,
),
),
MethodSpec(
"hyper_projection",
"southern_women",
"bipartite",
lambda c: nb.fraction_filter(
nb.hyper_projection(c["davis_bipartite"], c["women_nodes"]),
"weight",
0.3,
ascending=False,
),
),
MethodSpec(
"probs_projection",
"southern_women",
"bipartite",
lambda c: nb.fraction_filter(
nb.probs_projection(c["davis_bipartite"], c["women_nodes"]),
"weight",
0.3,
ascending=False,
),
),
MethodSpec(
"ycn_projection",
"southern_women",
"bipartite",
lambda c: nb.fraction_filter(
nb.ycn_projection(c["davis_bipartite"], c["women_nodes"]),
"weight",
0.3,
ascending=False,
),
),
MethodSpec(
"bipartite_projection",
"southern_women",
"bipartite",
lambda c: nb.fraction_filter(
nb.bipartite_projection(
c["davis_bipartite"],
c["women_nodes"],
method="simple",
),
"weight",
0.3,
ascending=False,
),
),
MethodSpec(
"sdsm",
"southern_women",
"bipartite",
lambda c: nb.threshold_filter(
nb.sdsm(c["davis_bipartite"], c["women_nodes"]),
"sdsm_pvalue",
0.05,
mode="below",
include_all_nodes=False,
),
),
MethodSpec(
"fdsm",
"southern_women",
"bipartite",
lambda c: nb.threshold_filter(
nb.fdsm(c["davis_bipartite"], c["women_nodes"], trials=250, seed=42),
"fdsm_pvalue",
0.05,
mode="below",
include_all_nodes=False,
),
),
MethodSpec(
"fixedfill",
"southern_women",
"bipartite",
lambda c: nb.fixedfill(c["davis_bipartite"], c["women_nodes"], alpha=0.05),
),
MethodSpec(
"fixedrow",
"southern_women",
"bipartite",
lambda c: nb.fixedrow(c["davis_bipartite"], c["women_nodes"], alpha=0.05),
),
MethodSpec(
"fixedcol",
"southern_women",
"bipartite",
lambda c: nb.fixedcol(c["davis_bipartite"], c["women_nodes"], alpha=0.05),
),
]
def _context() -> dict:
les_weighted = nx.les_miserables_graph()
les_unweighted = _to_unweighted(les_weighted)
davis = nx.davis_southern_women_graph()
women_nodes = [n for n, d in davis.nodes(data=True) if d.get("bipartite") == 0]
women_projection = nx.Graph()
women_projection.add_nodes_from(women_nodes)
for i, u in enumerate(women_nodes):
for v in women_nodes[i + 1 :]:
women_projection.add_edge(u, v)
return {
"les_weighted": les_weighted,
"les_unweighted": les_unweighted,
"davis_bipartite": davis,
"women_nodes": women_nodes,
"women_projection": women_projection,
}
def _base_graphs(ctx: dict) -> dict:
return {
"les_miserables": ctx["les_weighted"],
"southern_women": ctx["women_projection"],
}
context = _context()
base_graphs = _base_graphs(context)
positions = {
"les_miserables": nx.spring_layout(
base_graphs["les_miserables"],
seed=42,
weight="weight",
),
"southern_women": nx.spring_layout(base_graphs["southern_women"], seed=42),
}
rows = []
for spec in _method_specs():
base_graph = base_graphs[spec.dataset]
backbone = spec.build(context)
if backbone.number_of_edges() == base_graph.number_of_edges():
warnings.warn(
(
f"{spec.name} ({spec.dataset}) returned the same number of edges "
"as the original graph. Re-test and validate threshold settings."
),
UserWarning,
)
backbone = _drop_isolates(backbone)
if not base_graph.is_directed() and backbone.is_directed():
backbone = _to_undirected_simple(backbone)
fig, ax, diff = compare_graphs(
base_graph,
backbone,
pos=positions[spec.dataset],
title=f"{_title(spec.name)} ({spec.dataset})",
return_diff=True,
)
rows.append(
{
"name": spec.name,
"module": spec.module,
"dataset": spec.dataset,
"original_edges": base_graph.number_of_edges(),
"backbone_edges": len(diff["kept_edges"]),
"removed_nodes": len(diff["removed_nodes"]),
}
)
print("Method | Dataset | Original Edges | Backbone Edges | Removed Nodes")
print("--- | --- | ---: | ---: | ---:")
for row in rows:
print(
f"{row['name']} | {row['dataset']} | {row['original_edges']} | "
f"{row['backbone_edges']} | {row['removed_nodes']}"
)
Total running time of the script: (0 minutes 10.934 seconds)