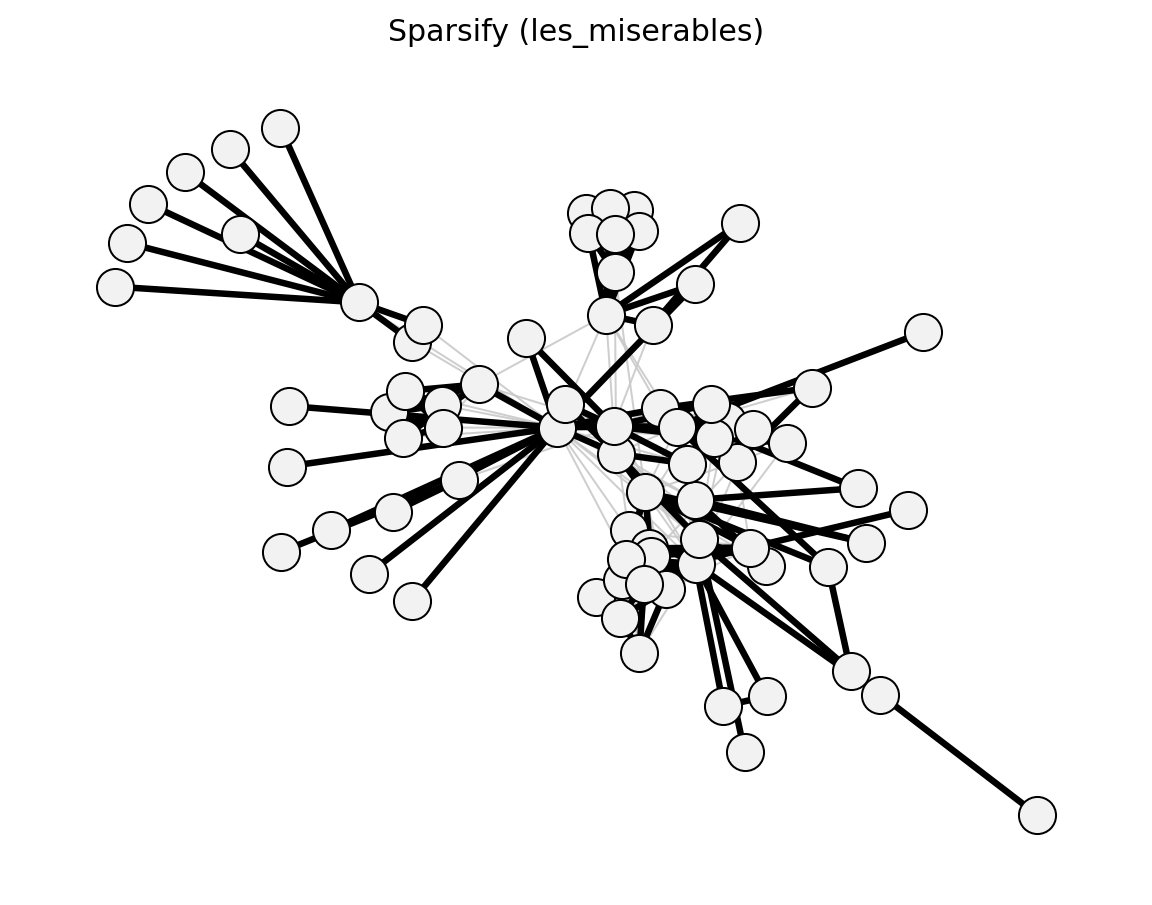
Unweighted Sparsification#
Examples in this module use the unweighted version of
nx.les_miserables_graph().
Methods return scored full graphs with sparsify_score and
sparsify_keep; apply boolean_filter() to
extract sparse backbones.
Complexity classes are provided in each function docstring.
Unweighted network backbone methods.
These methods sparsify unweighted graphs by scoring edges and filtering based on local topology.
- networkx_backbone.unweighted.sparsify()[source]#
Generic sparsification framework (score -> normalise -> filter -> connect).
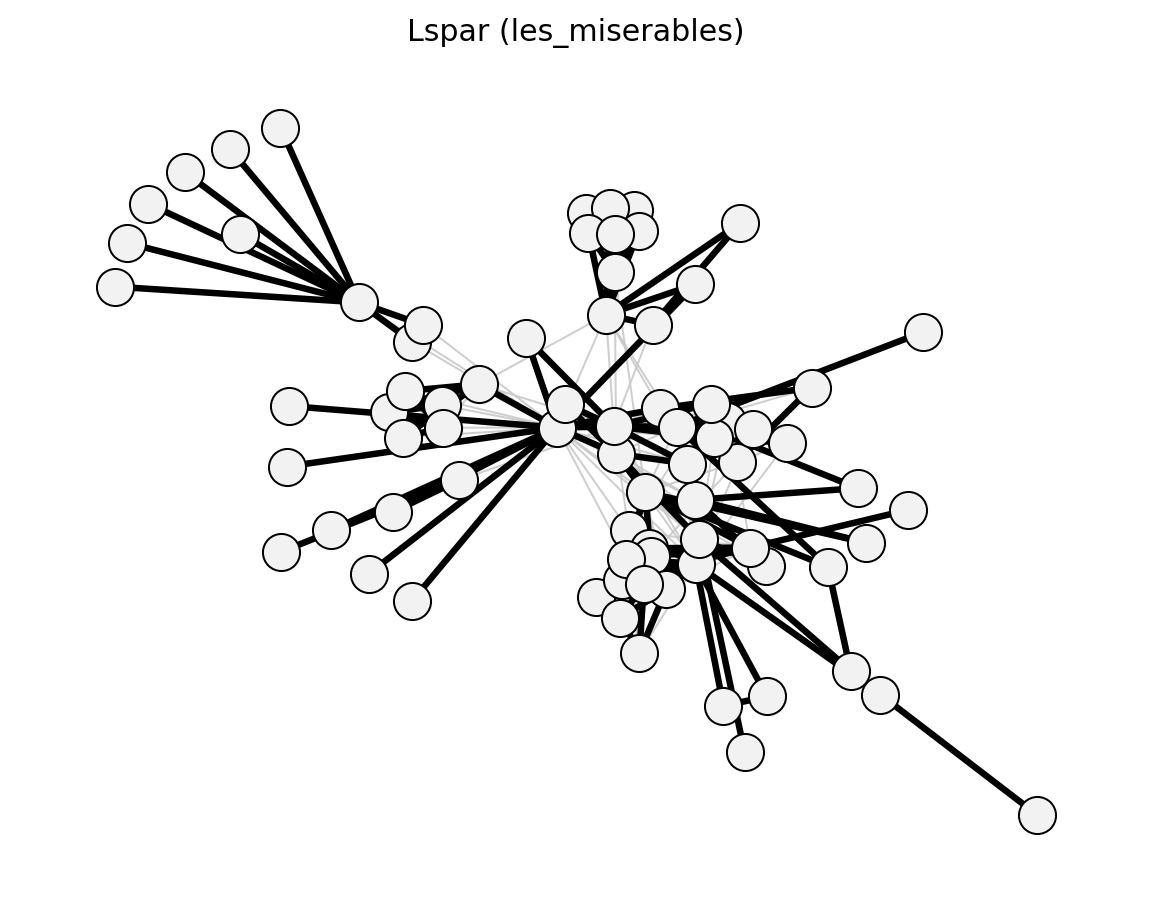
- networkx_backbone.unweighted.lspar()[source]#
Local Sparsification (Satuluri et al. 2011) – Jaccard-based.
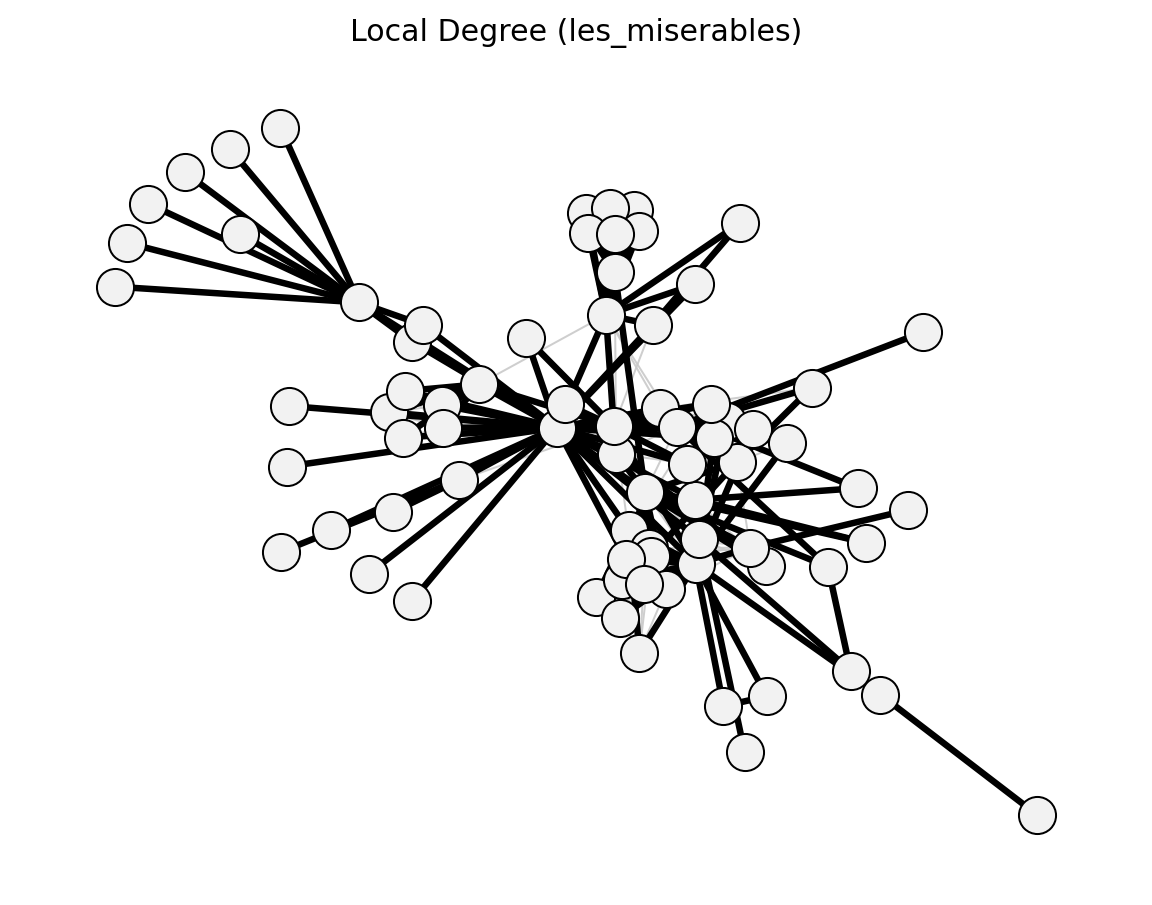
- networkx_backbone.unweighted.local_degree()[source]#
Local Degree model (Hamann et al. 2016) – degree-based scoring.
- networkx_backbone.sparsify(G, escore='jaccard', normalize='rank', filter='degree', s=0.5, umst=False)[source]#
Sparsify an unweighted graph using a score-normalise-filter pipeline.
Follows the four-step pipeline from Neal (2022) [1]:
Score: Assign a relevance score to each edge.
Normalise: Optionally rank-transform scores within each node’s neighbourhood.
Filter: Retain a fraction of edges per node or globally.
Connect: Optionally add the union of minimum spanning trees to ensure connectivity.
- Parameters:
G (graph) – A NetworkX graph.
escore (string, optional (default="jaccard")) – Edge scoring method. One of
"jaccard","degree","triangles","quadrangles", or"random".normalize (string or None, optional (default="rank")) – Normalisation method.
"rank"normalises scores by ranking within each node’s neighbourhood.Noneskips normalisation.filter (string, optional (default="degree")) – Filtering method.
"degree"keepsceil(d^s)top-scored edges per node."threshold"keeps edges with normalised score >= s.s (float, optional (default=0.5)) – Sparsification parameter.
umst (bool, optional (default=False)) – If
True, add the union of minimum spanning trees to guarantee connectivity.
- Returns:
H – A copy of G with edge attributes:
"sparsify_score"and boolean"sparsify_keep".- Return type:
graph
- Raises:
ValueError – If filter is not
"degree"or"threshold".
References
[1]Neal, Z. P. (2022). backbone: An R package to extract network backbones. PLOS ONE, 17(5), e0269137.
Examples
>>> import networkx as nx >>> from networkx_backbone import boolean_filter, sparsify >>> G_weighted = nx.les_miserables_graph() >>> G = nx.Graph() >>> G.add_nodes_from(G_weighted.nodes(data=True)) >>> G.add_edges_from(G_weighted.edges()) >>> H = sparsify(G, s=0.5) >>> backbone = boolean_filter(H, "sparsify_keep") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(mn)time andO(n + m)space.
- networkx_backbone.lspar(G, s=0.5)[source]#
Compute the Local Sparsification backbone.
Convenience wrapper around
sparsify()using Jaccard scoring, rank normalisation, and degree filtering [1].- Parameters:
G (graph) – A NetworkX graph.
s (float, optional (default=0.5)) – Sparsification exponent (0 = sparsest, 1 = keep all).
- Returns:
H – A copy of G with
"sparsify_score"and"sparsify_keep".- Return type:
graph
References
[1]Satuluri, V., Parthasarathy, S., & Ruan, Y. (2011). Local graph sparsification for scalable clustering. ACM SIGMOD, 721-732.
Examples
>>> import networkx as nx >>> from networkx_backbone import boolean_filter, lspar >>> G_weighted = nx.les_miserables_graph() >>> G = nx.Graph() >>> G.add_nodes_from(G_weighted.nodes(data=True)) >>> G.add_edges_from(G_weighted.edges()) >>> H = lspar(G, s=0.5) >>> backbone = boolean_filter(H, "sparsify_keep") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(mn)time andO(n + m)space.
- networkx_backbone.local_degree(G, s=0.3)[source]#
Compute the Local Degree backbone.
Convenience wrapper around
sparsify()using degree scoring, rank normalisation, and degree filtering [1].- Parameters:
G (graph) – A NetworkX graph.
s (float, optional (default=0.3)) – Sparsification exponent (0 = sparsest, 1 = keep all).
- Returns:
H – A copy of G with
"sparsify_score"and"sparsify_keep".- Return type:
graph
References
[1]Hamann, M., Lindner, G., Meyerhenke, H., Staudt, C. L., & Wagner, D. (2016). Structure-preserving sparsification methods for social networks. Social Network Analysis and Mining, 6(1), 22.
Examples
>>> import networkx as nx >>> from networkx_backbone import boolean_filter, local_degree >>> G_weighted = nx.les_miserables_graph() >>> G = nx.Graph() >>> G.add_nodes_from(G_weighted.nodes(data=True)) >>> G.add_edges_from(G_weighted.edges()) >>> H = local_degree(G, s=0.3) >>> backbone = boolean_filter(H, "sparsify_keep") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(m + n)time andO(n + m)space.
Gallery Examples#
Function Image Reference