Structural Methods#
Examples in this module use nx.les_miserables_graph().
These functions return scored full graphs. Extract final backbones with
boolean_filter() on *_keep flags (or
threshold_filter() for continuous scores such as
salience and ds_weight).
Complexity classes are provided in each function docstring.
Structural backbone extraction methods.
These methods use the network’s topology (edge weights, shortest paths, spanning trees, degree, neighborhood overlap) to identify the most important substructure.
- networkx_backbone.structural.global_sparsification()[source]#
Keep top-weight edges globally by fraction.
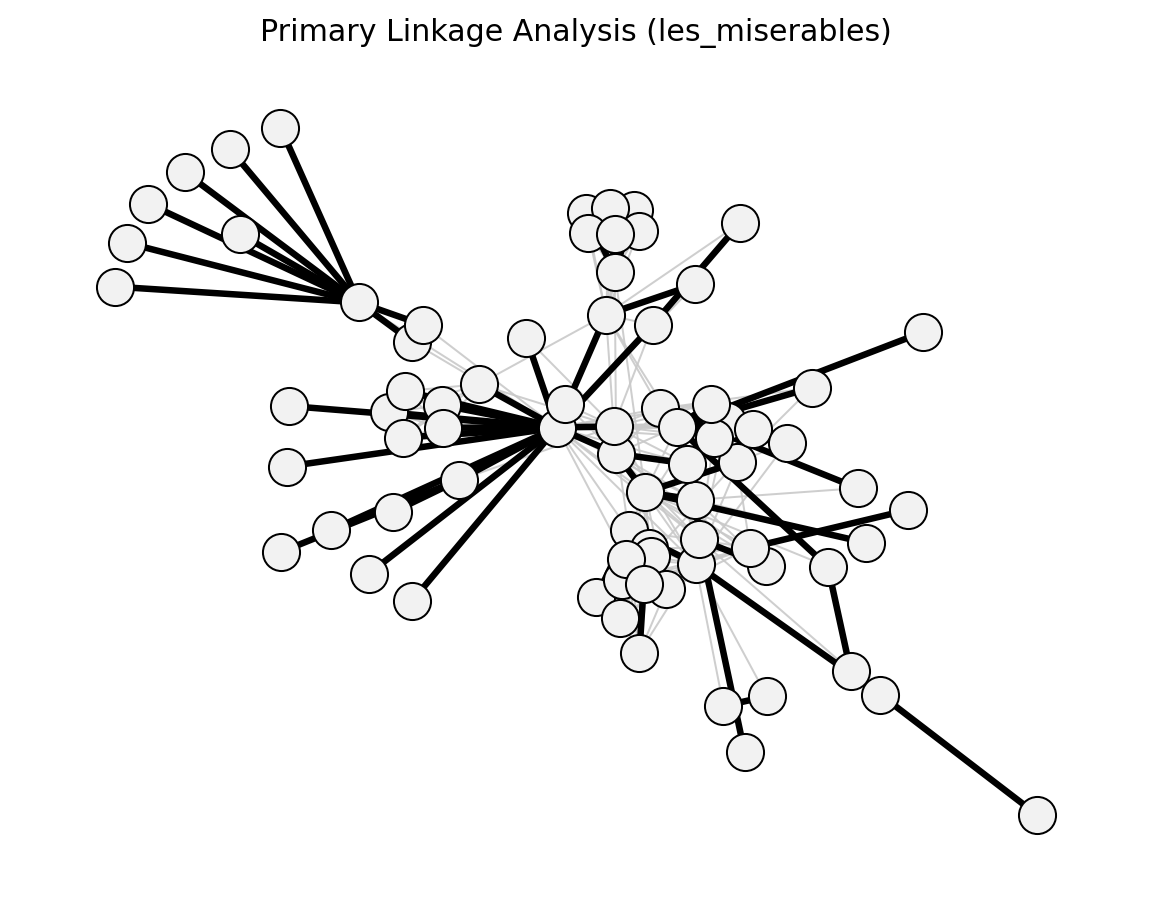
- networkx_backbone.structural.primary_linkage_analysis()[source]#
Nystuen & Dacey (1961) – strongest outgoing edge per node.
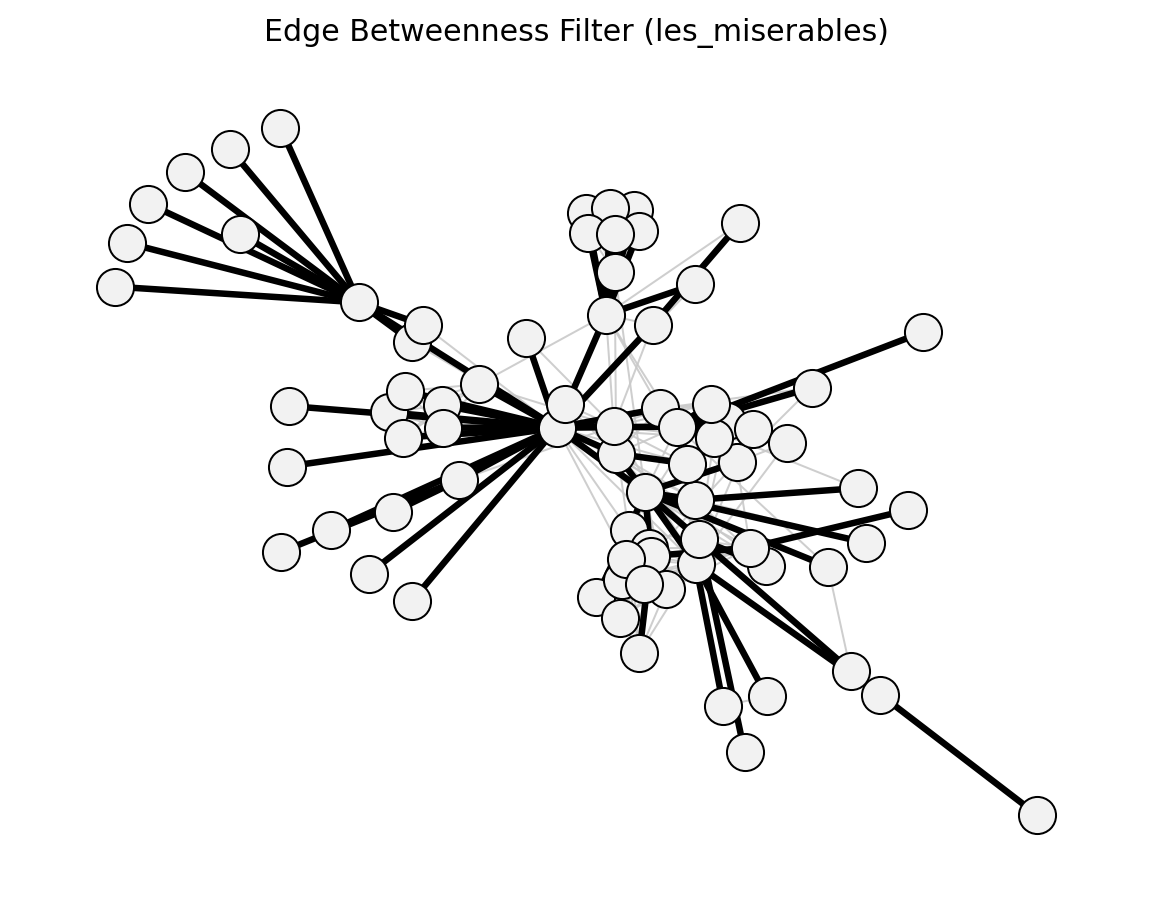
- networkx_backbone.structural.edge_betweenness_filter()[source]#
Girvan & Newman (2002) – keep top edge-betweenness edges.
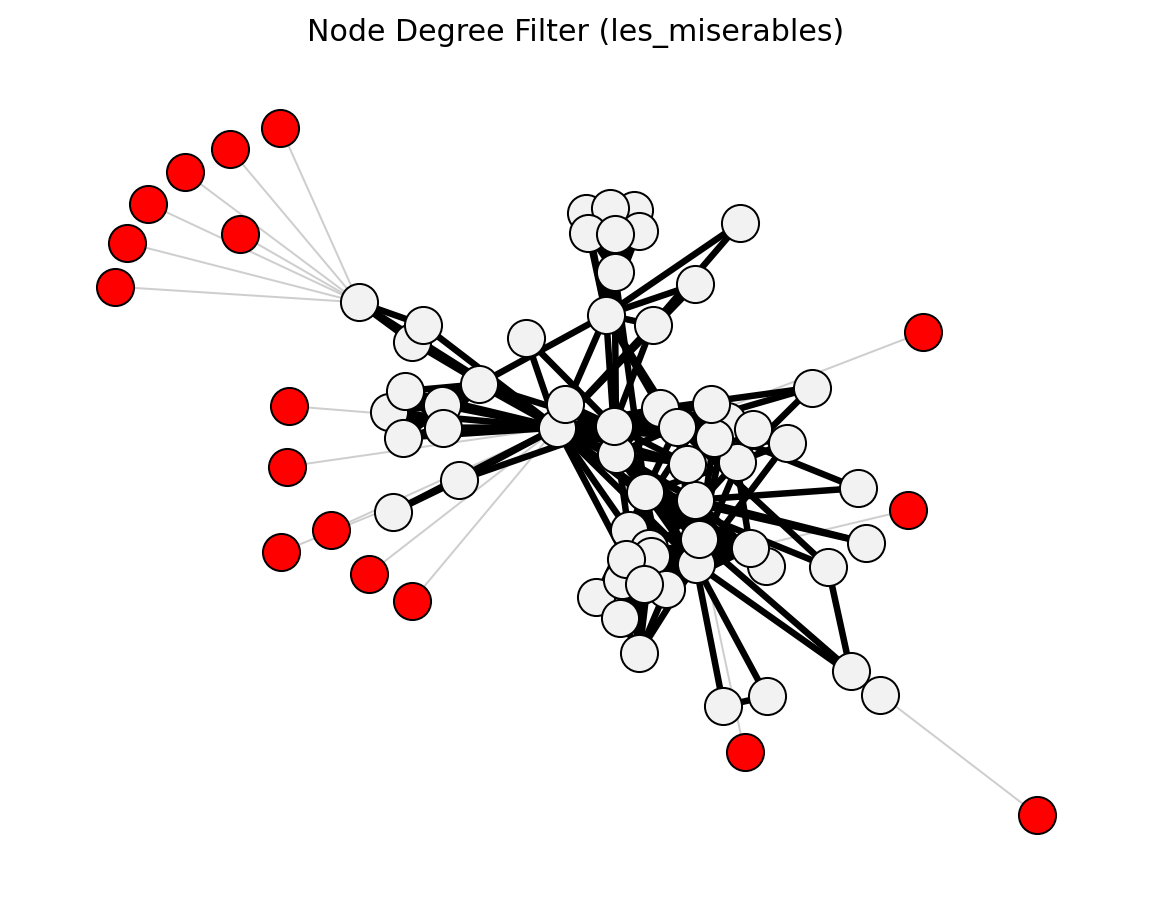
- networkx_backbone.structural.node_degree_filter()[source]#
Keep nodes above a minimum degree threshold.
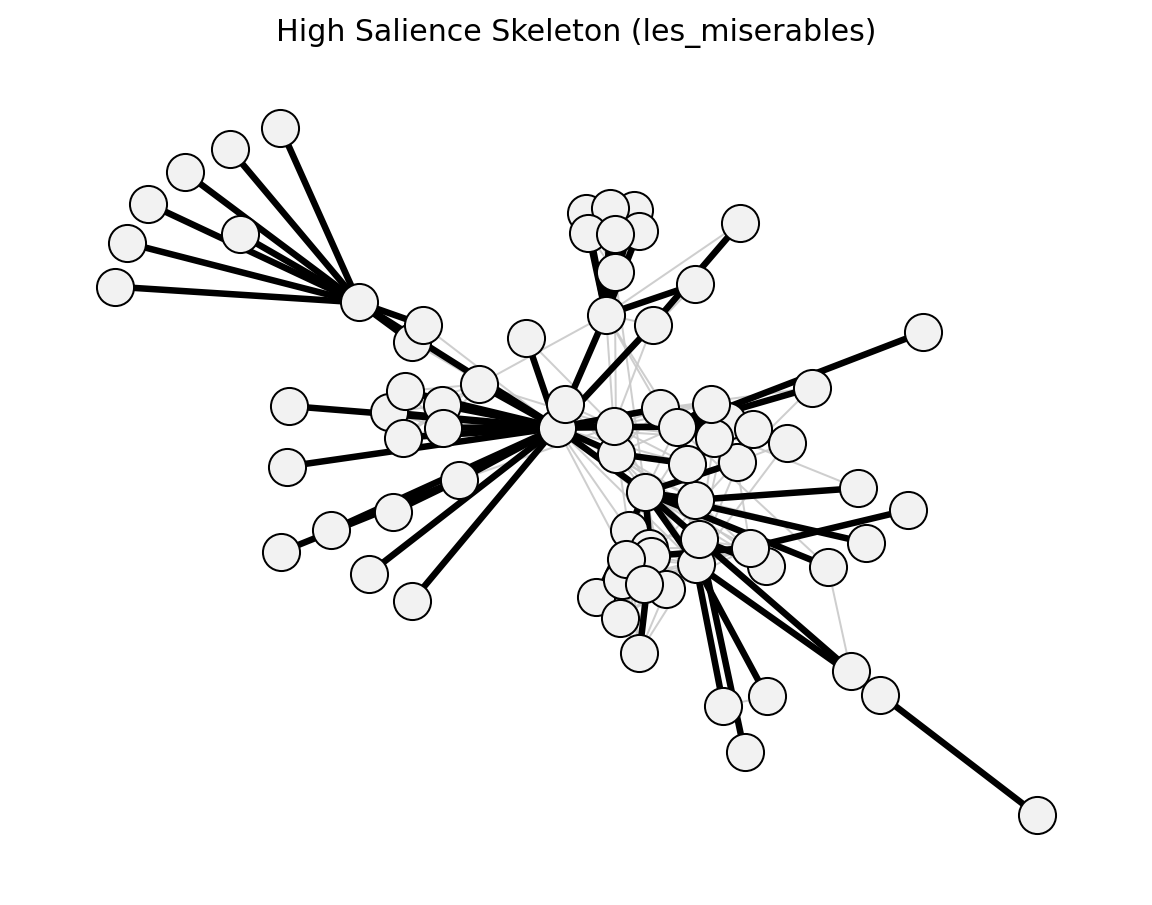
- networkx_backbone.structural.high_salience_skeleton()[source]#
Grady et al. (2012) – shortest-path tree participation.
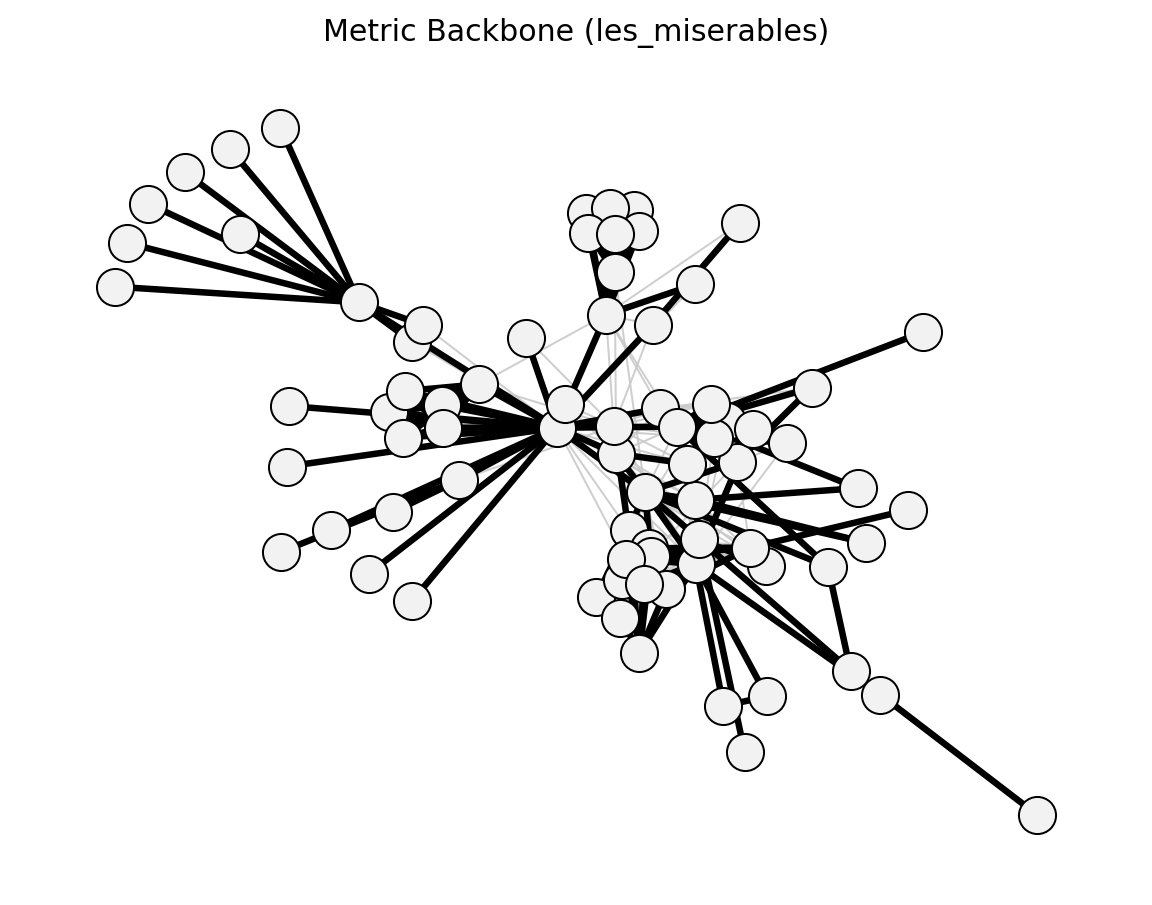
- networkx_backbone.structural.metric_backbone()[source]#
Simas et al. (2021) – edges on shortest paths (sum distances).
- networkx_backbone.structural.doubly_stochastic_filter()[source]#
Slater (2009) – Sinkhorn-Knopp normalization.
- networkx_backbone.structural.modularity_backbone()[source]#
Rajeh et al. (2022) – node vitality index.
- networkx_backbone.structural.planar_maximally_filtered_graph()[source]#
Tumminello et al. (2005) – planar constraint.
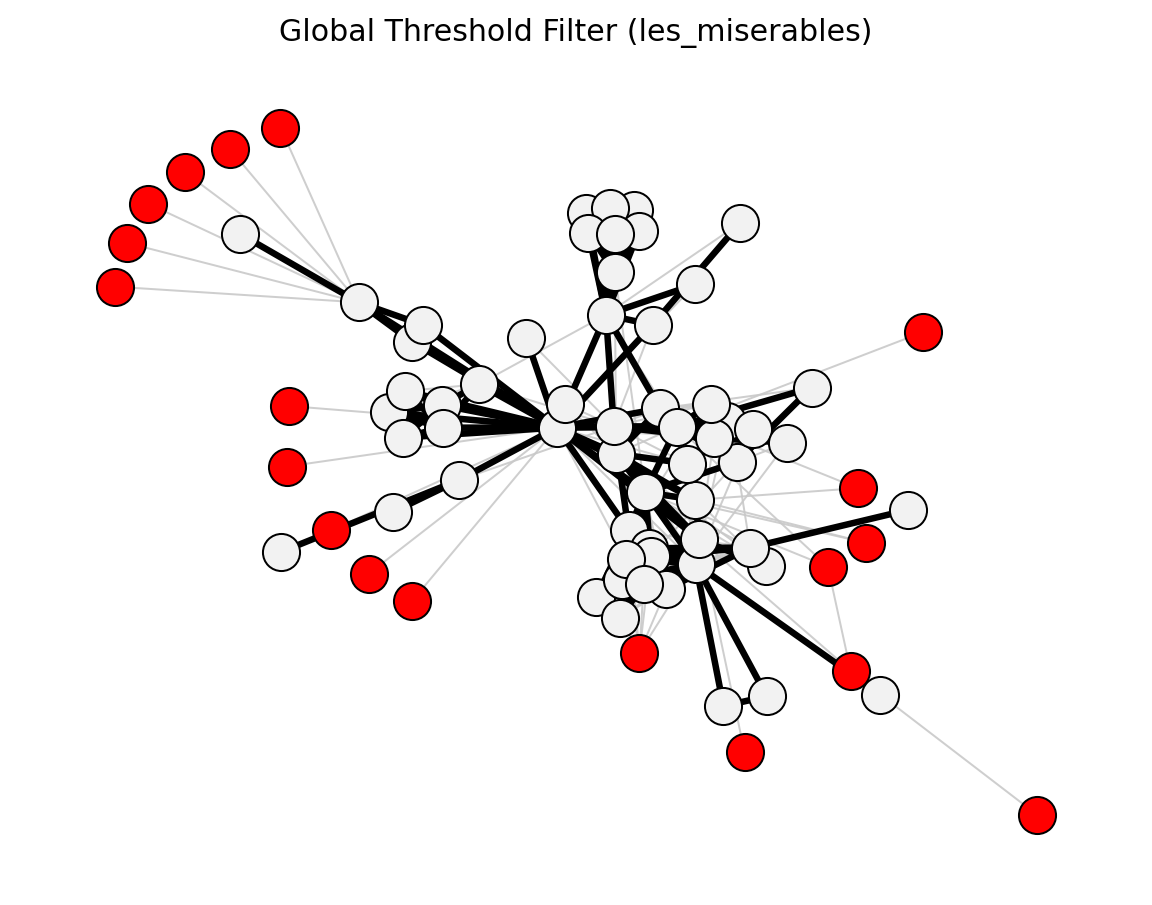
- networkx_backbone.global_threshold_filter(G, threshold, weight='weight')[source]#
Score edges against a global weight threshold.
For each edge, mark whether its weight attribute is greater than or equal to threshold.
- Parameters:
G (graph) – A NetworkX graph.
threshold (float) – Minimum edge weight required. Edges with
data[weight] >= thresholdare kept.weight (string, optional (default="weight")) – Edge attribute key used as the weight.
- Returns:
H – A copy of G with a boolean
"global_threshold_keep"edge attribute.- Return type:
graph
Examples
>>> import networkx as nx >>> from networkx_backbone import boolean_filter, global_threshold_filter >>> G = nx.les_miserables_graph() >>> H = global_threshold_filter(G, threshold=3.0) >>> backbone = boolean_filter(H, "global_threshold_keep") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(m)time andO(n + m)space.
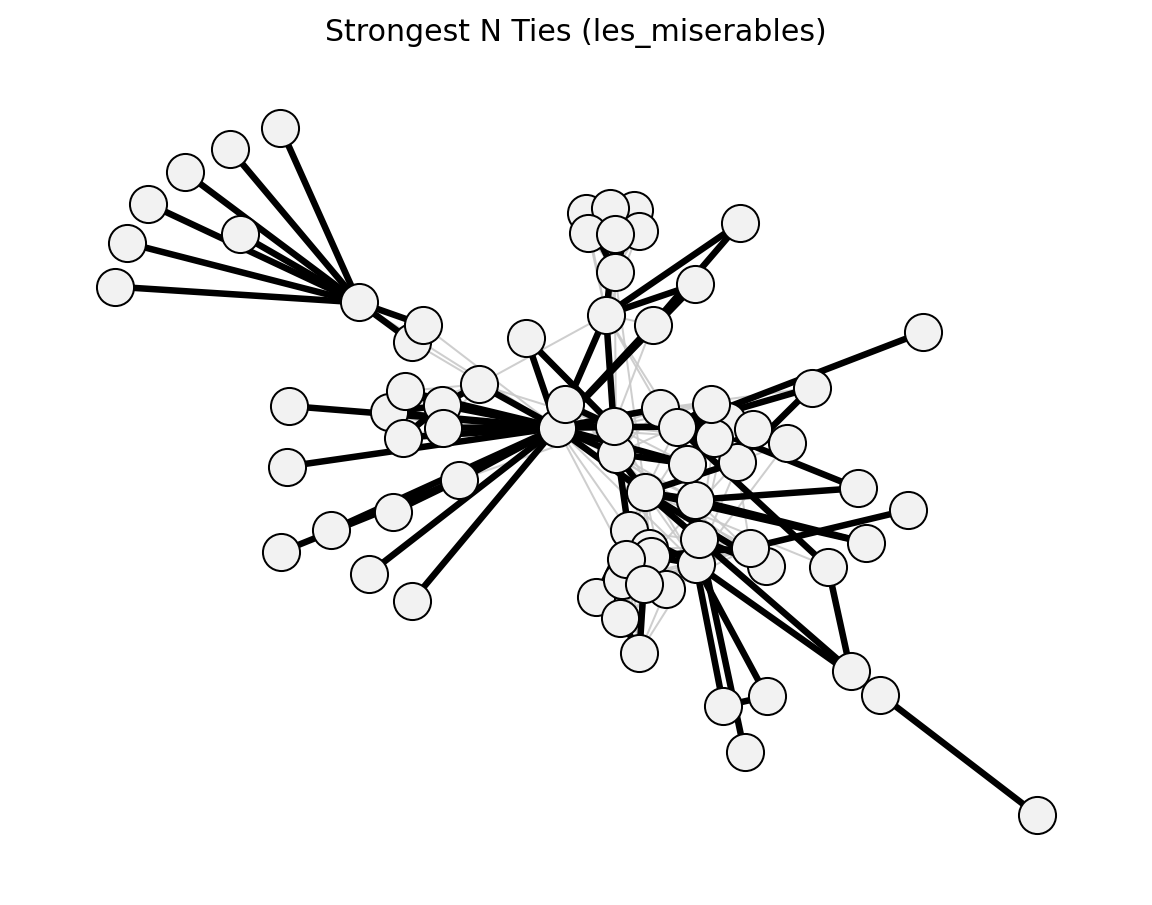
- networkx_backbone.strongest_n_ties(G, n=1, weight='weight')[source]#
Score edges by strongest-n membership per node.
For each node, select the n incident edges with the largest weight value. An edge is kept if either endpoint selects it (OR semantics). For directed graphs the selection is based on out-edges.
- Parameters:
G (graph) – A NetworkX graph.
n (int, optional (default=1)) – Number of strongest edges to keep per node. Must be at least 1.
weight (string, optional (default="weight")) – Edge attribute key used as the weight.
- Returns:
H – A copy of G with a boolean
"strongest_n_ties_keep"edge attribute.- Return type:
graph
- Raises:
ValueError – If n < 1.
Examples
>>> import networkx as nx >>> from networkx_backbone import boolean_filter, strongest_n_ties >>> G = nx.les_miserables_graph() >>> H = strongest_n_ties(G, n=1) >>> backbone = boolean_filter(H, "strongest_n_ties_keep") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(m log n)time andO(n + m)space.
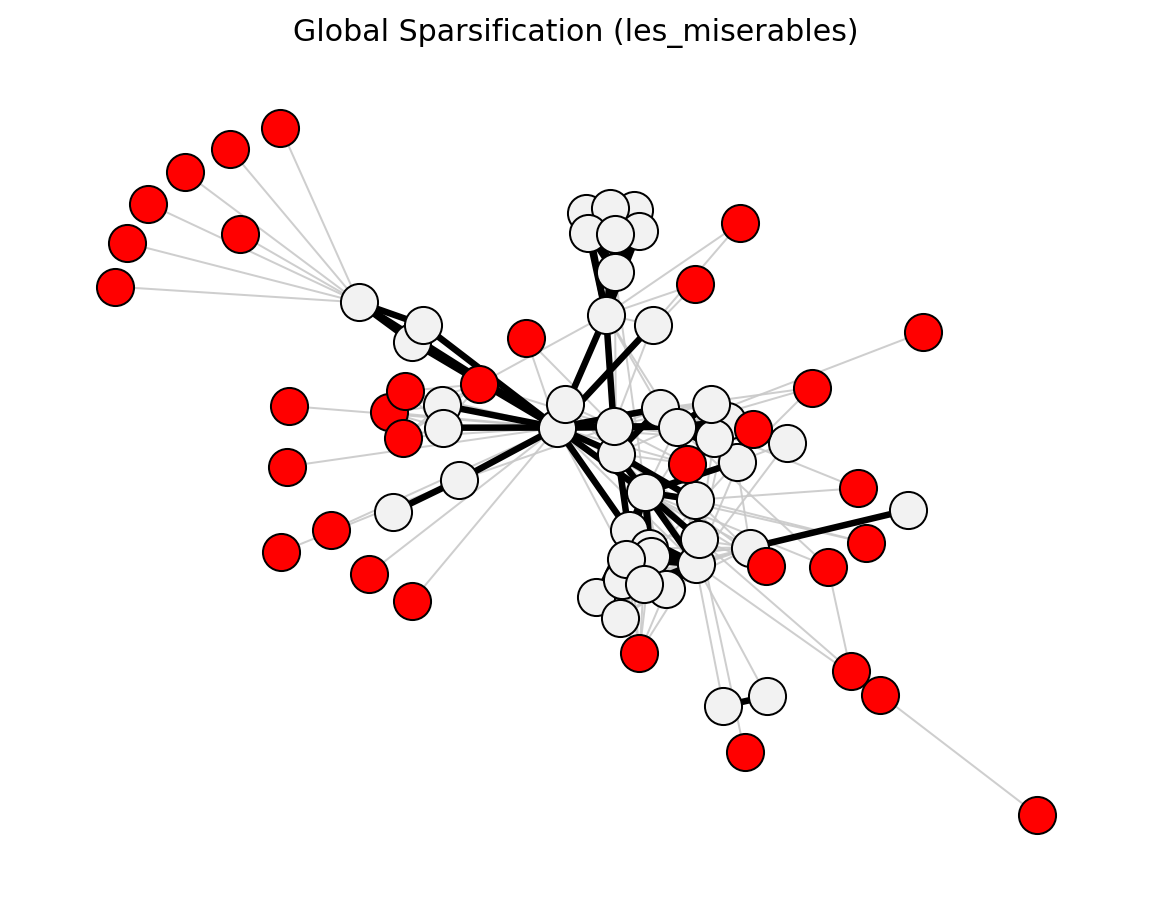
- networkx_backbone.global_sparsification(G, s=0.5, weight='weight')[source]#
Score edges by global top-fraction membership.
This method ranks all edges by weight and keeps the top
sfraction.- Parameters:
G (graph) – A NetworkX graph.
s (float, optional (default=0.5)) – Fraction of edges to retain. Must be in (0, 1].
weight (string, optional (default="weight")) – Edge attribute key used as the weight.
- Returns:
H – A copy of G with a boolean
"global_sparsification_keep"edge attribute.- Return type:
graph
- Raises:
ValueError – If
sis outside (0, 1].
References
[1]Satuluri, V., Parthasarathy, S., & Ruan, Y. (2011). Local graph sparsification for scalable clustering. SIGMOD, 721-732.
Computational complexity estimate is
O(m log m)time andO(n + m)space.
- networkx_backbone.primary_linkage_analysis(G, weight='weight')[source]#
Score edges by primary-linkage membership.
Each node keeps only its strongest outgoing connection.
- Parameters:
G (networkx.Graph or networkx.DiGraph) – Input weighted graph.
weight (string, optional (default="weight")) – Edge attribute key used as the weight.
- Returns:
H – A copy of G with a boolean
"primary_linkage_keep"edge attribute.- Return type:
graph
References
[1]Nystuen, J. D., & Dacey, M. F. (1961). A graph theory interpretation of nodal regions. Papers in Regional Science, 7(1), 29-42.
Computational complexity estimate is
O(m)time andO(n + m)space.
- networkx_backbone.edge_betweenness_filter(G, s=0.5, weight='weight')[source]#
Score edges by edge betweenness and top-fraction membership.
- Parameters:
G (graph) – A NetworkX graph.
s (float, optional (default=0.5)) – Fraction of edges to retain. Must be in (0, 1].
weight (string, optional (default="weight")) – Edge attribute key used as the weight. If present, distance is interpreted as
1 / weight.
- Returns:
H – A copy of G with
"edge_betweenness"scores and a boolean"edge_betweenness_keep"edge attribute.- Return type:
graph
- Raises:
ValueError – If
sis outside (0, 1].
References
[1]Girvan, M., & Newman, M. E. J. (2002). Community structure in social and biological networks. PNAS, 99(12), 7821-7826.
Computational complexity estimate is
O(nm + n^2 log n)time andO(n + m)space.
- networkx_backbone.node_degree_filter(G, min_degree=1)[source]#
Score nodes and edges by a minimum node-degree criterion.
- Parameters:
G (graph) – A NetworkX graph.
min_degree (int, optional (default=1)) – Minimum degree required for a node to be retained.
- Returns:
H – A copy of G with boolean
"node_degree_keep"on nodes and edges.- Return type:
graph
- Raises:
ValueError – If
min_degreeis negative.
References
[1]Freeman, L. C. (1978). Centrality in social networks: conceptual clarification. Social Networks, 1(3), 215-239.
Computational complexity estimate is
O(n + m)time andO(n + m)space.
- networkx_backbone.high_salience_skeleton(G, weight='weight')[source]#
Compute edge salience from shortest-path tree participation.
For every node r, a shortest-path tree rooted at r is computed (using inverse weights as distances). The salience of an edge is the fraction of these trees that contain it. The salience is stored as the
"salience"edge attribute on the returned graph.- Parameters:
G (graph) – A NetworkX graph. All weights must be positive.
weight (string, optional (default="weight")) – Edge attribute key used as the weight.
- Returns:
H – A copy of G with an additional
"salience"edge attribute whose value lies in [0, 1].- Return type:
graph
References
[1]Grady, D., Thiemann, C., & Brockmann, D. (2012). Robust classification of salient links in complex networks. Nature Communications, 3, 864.
Examples
>>> import networkx as nx >>> from networkx_backbone import high_salience_skeleton, threshold_filter >>> G = nx.les_miserables_graph() >>> H = high_salience_skeleton(G) >>> backbone = threshold_filter(H, "salience", 0.5, mode="above") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(nm + n^2 log n)time andO(n + m)space.
- networkx_backbone.metric_backbone(G, weight='weight')[source]#
Extract the metric backbone using sum-of-distances shortest paths.
An edge (u, v) is in the metric backbone if and only if its direct distance (the inverse of its weight) equals the shortest-path distance between u and v computed over the full graph.
- Parameters:
G (graph) – A NetworkX graph.
weight (string, optional (default="weight")) – Edge attribute key used as the weight. All weights must be positive.
- Returns:
H – A copy of G with boolean
"metric_keep"on each edge.- Return type:
graph
References
[1]Simas, T., Correia, R. B., & Rocha, L. M. (2021). The distance backbone of complex networks. J. Complex Netw., 9, cnab021.
Examples
>>> import networkx as nx >>> from networkx_backbone import boolean_filter, metric_backbone >>> G = nx.les_miserables_graph() >>> H = metric_backbone(G) >>> backbone = boolean_filter(H, "metric_keep") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(nm + n^2 log n)time andO(n^2 + m)space.
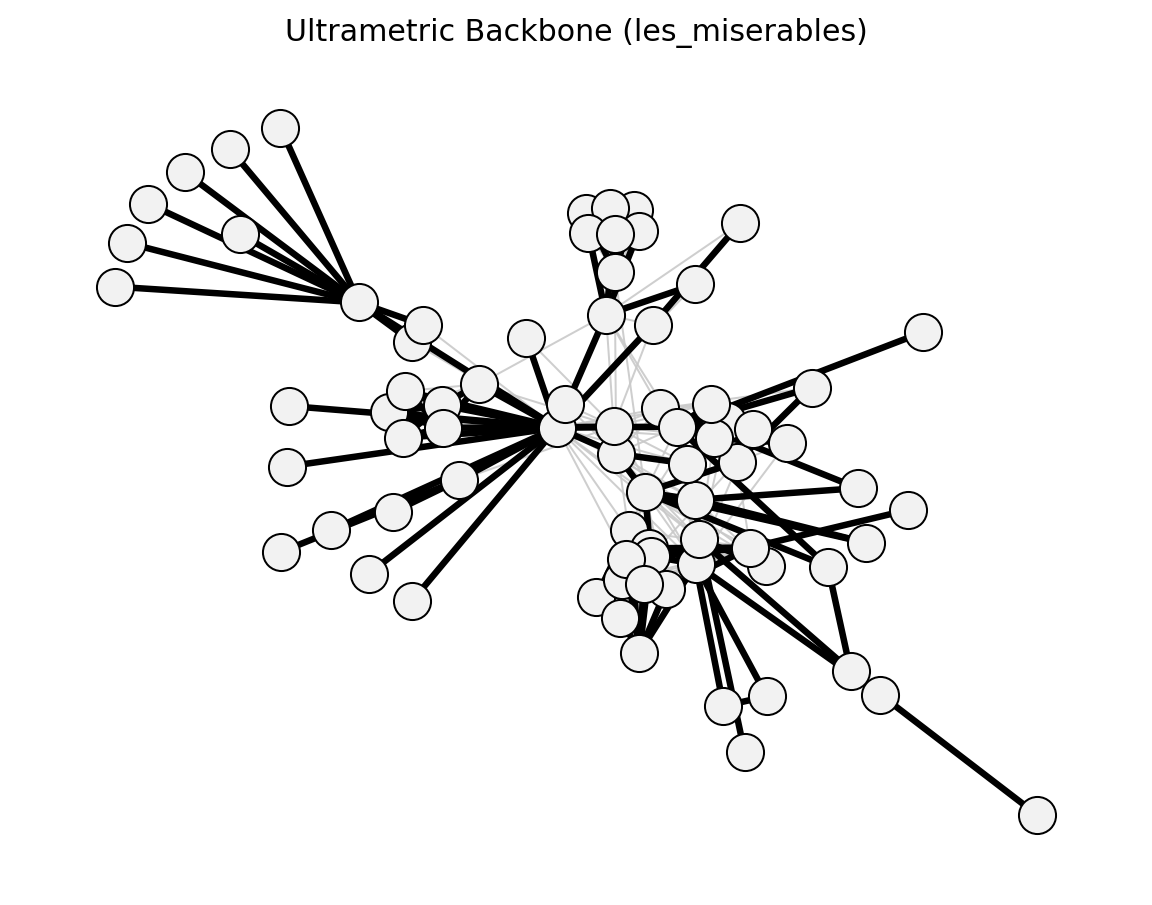
- networkx_backbone.ultrametric_backbone(G, weight='weight')[source]#
Extract the ultrametric backbone using max-distance (minimax) paths.
An edge (u, v) is in the ultrametric backbone if its direct distance is no greater than the minimax path distance between u and v (the path that minimises the maximum edge distance along the path).
- Parameters:
G (graph) – A NetworkX graph.
weight (string, optional (default="weight")) – Edge attribute key used as the weight. All weights must be positive.
- Returns:
H – A copy of G with boolean
"ultrametric_keep"on each edge.- Return type:
graph
References
[1]Simas, T., Correia, R. B., & Rocha, L. M. (2021). The distance backbone of complex networks. J. Complex Netw., 9, cnab021.
Examples
>>> import networkx as nx >>> from networkx_backbone import boolean_filter, ultrametric_backbone >>> G = nx.les_miserables_graph() >>> H = ultrametric_backbone(G) >>> backbone = boolean_filter(H, "ultrametric_keep") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(mn + m log n)time andO(n + m)space.
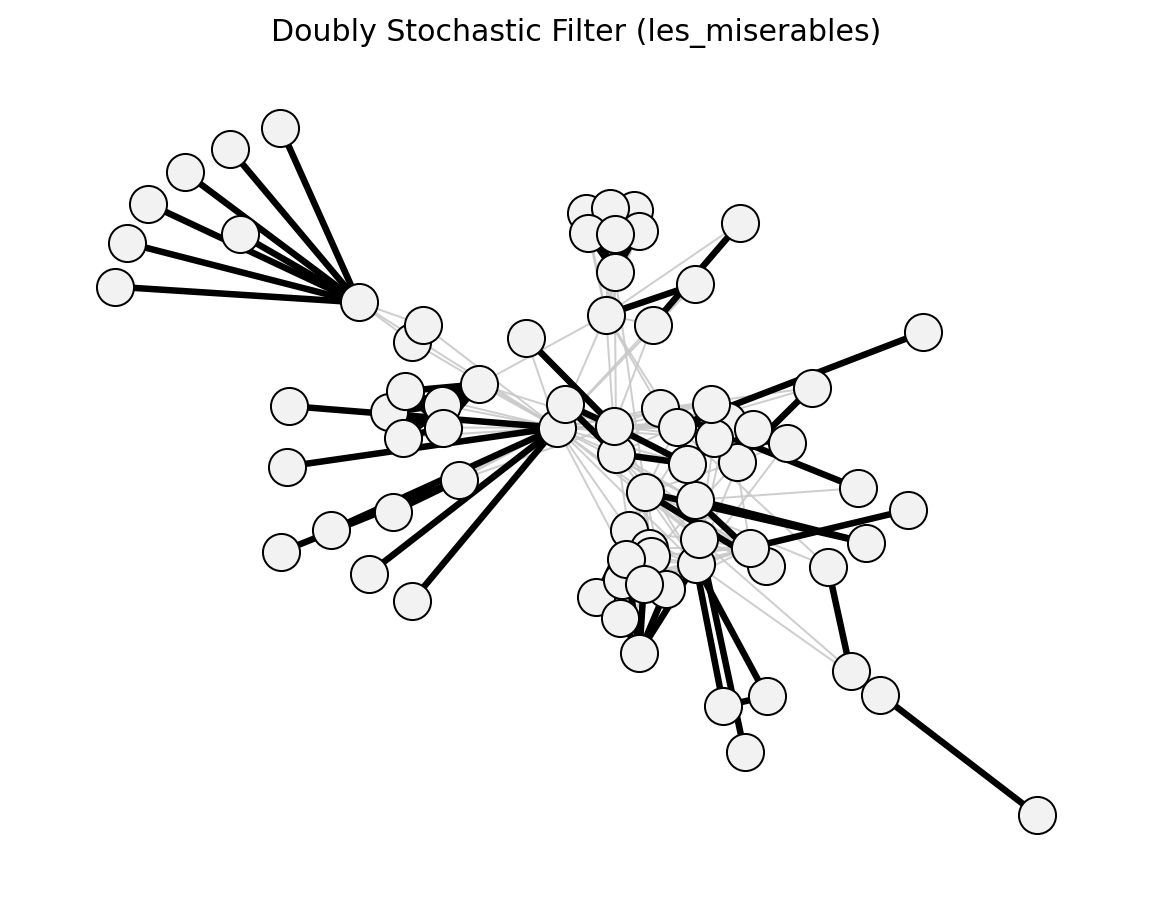
- networkx_backbone.doubly_stochastic_filter(G, weight='weight', max_iter=1000, tol=1e-08)[source]#
Compute the doubly-stochastic backbone via Sinkhorn-Knopp normalization.
Transforms the adjacency matrix into a doubly stochastic matrix via iterative Sinkhorn-Knopp normalization, then stores the normalised weight as the
"ds_weight"edge attribute on the returned graph.- Parameters:
- Returns:
H – A copy of G with an additional
"ds_weight"edge attribute (doubly-stochastic normalised weight).- Return type:
graph
References
[1]Slater, P. B. (2009). A two-stage algorithm for extracting the multiscale backbone of complex weighted networks. PNAS, 106(26), E66.
Examples
>>> import networkx as nx >>> from networkx_backbone import doubly_stochastic_filter, threshold_filter >>> G = nx.les_miserables_graph() >>> H = doubly_stochastic_filter(G) >>> backbone = threshold_filter(H, "ds_weight", 0.1, mode="above") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(I * n^2 + m)time andO(n^2 + m)space.
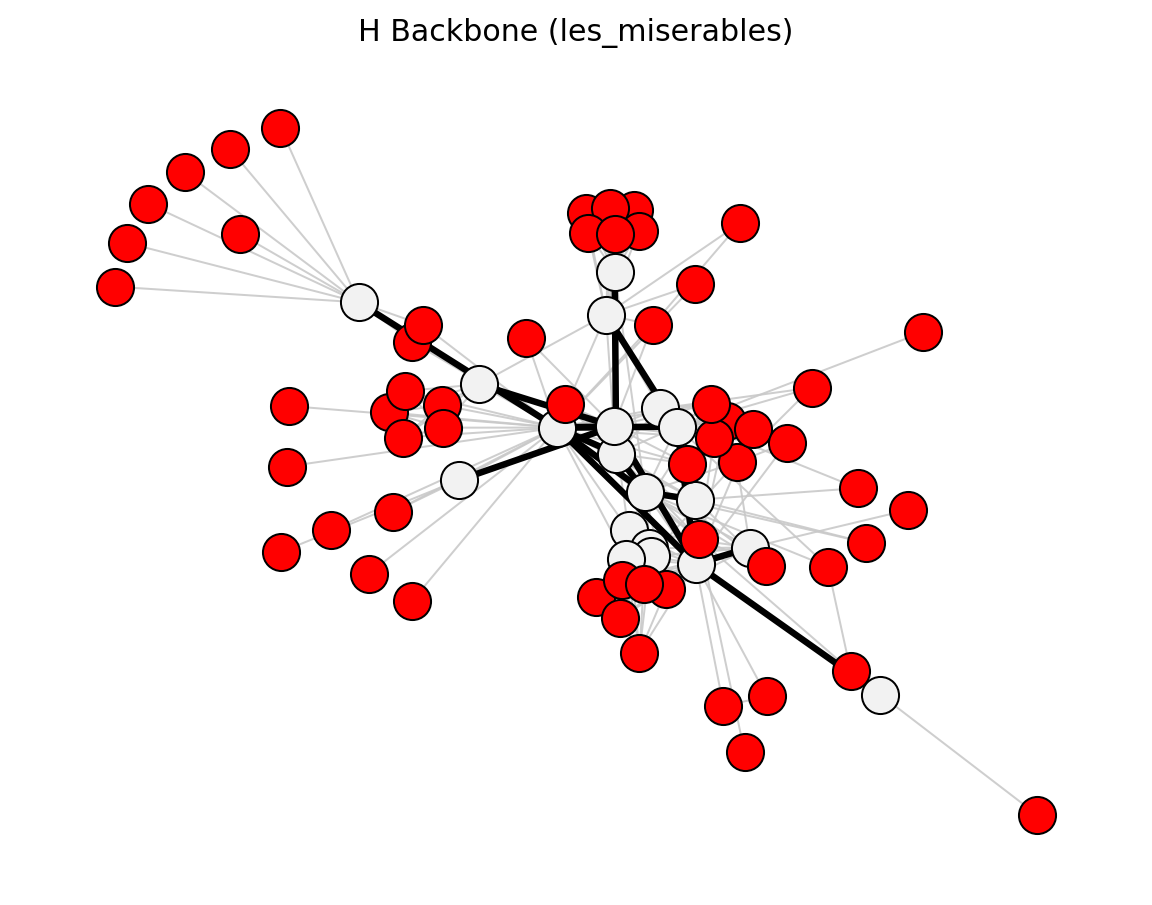
- networkx_backbone.h_backbone(G, weight='weight')[source]#
Extract the h-backbone of a weighted graph.
The h-backbone [1] is composed of two parts:
h-strength network: find h such that there are at least h edges with weight >= h. Keep those edges.
h-bridge network: among the remaining edges, keep those whose edge betweenness centrality is in the top-h.
The union of both subsets forms the h-backbone.
- Parameters:
G (graph) – A NetworkX graph.
weight (string, optional (default="weight")) – Edge attribute key used as the weight.
- Returns:
H – A copy of G with boolean
"h_backbone_keep"on each edge.- Return type:
graph
References
[1]Zhang, R. J., Stanley, H. E., & Ye, F. Y. (2018). Extracting h-backbone as a core structure in weighted networks. Scientific Reports, 8, 14356.
Examples
>>> import networkx as nx >>> from networkx_backbone import boolean_filter, h_backbone >>> G = nx.les_miserables_graph() >>> H = h_backbone(G) >>> backbone = boolean_filter(H, "h_backbone_keep") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(nm + n^2 log n + m log m)time andO(n + m)space.
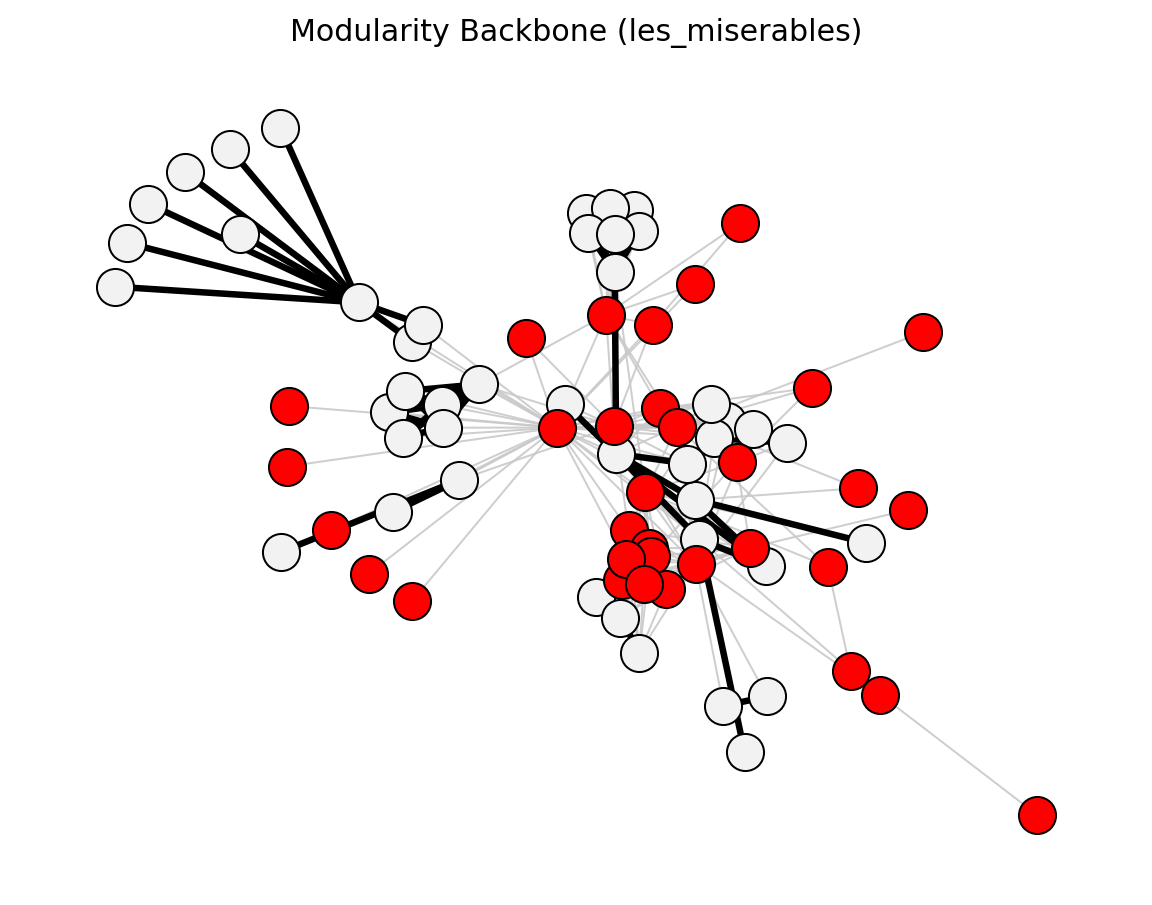
- networkx_backbone.modularity_backbone(G, weight='weight')[source]#
Compute the modularity vitality index for each node.
The vitality index [1] measures the change in modularity when a node is removed. Nodes that contribute positively to community structure have positive vitality. The vitality is stored as the
"vitality"node attribute on the returned graph.- Parameters:
G (graph) – A NetworkX graph.
weight (string, optional (default="weight")) – Edge attribute key used as the weight.
- Returns:
H – A copy of G with
"vitality"node scores and boolean"modularity_keep"on edges.- Return type:
graph
References
[1]Rajeh, S., Savonnet, M., Leclercq, E., & Cherifi, H. (2022). Modularity-based backbone extraction in weighted complex networks. NetSci-X 2022, 67-79.
Examples
>>> import networkx as nx >>> from networkx_backbone import boolean_filter, modularity_backbone >>> G = nx.les_miserables_graph() >>> H = modularity_backbone(G) >>> backbone = boolean_filter(H, "modularity_keep") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(nm)time andO(n + m)space.
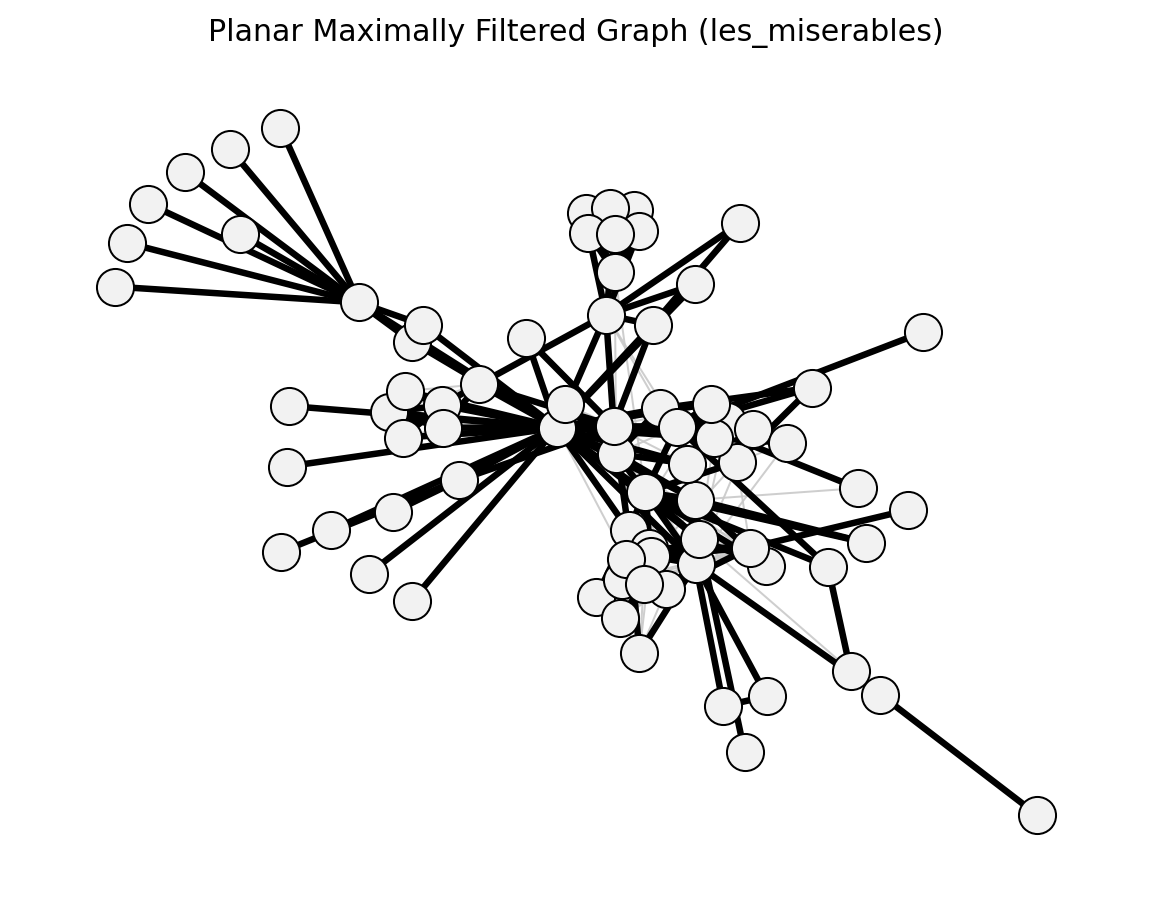
- networkx_backbone.planar_maximally_filtered_graph(G, weight='weight')[source]#
Construct the Planar Maximally Filtered Graph (PMFG).
Iteratively adds edges from heaviest to lightest, keeping only those whose addition preserves planarity [1].
- Parameters:
G (graph) – A NetworkX graph.
weight (string, optional (default="weight")) – Edge attribute key used as the weight.
- Returns:
H – A copy of G with boolean
"pmfg_keep"on edges.- Return type:
graph
References
[1]Tumminello, M., Aste, T., Di Matteo, T., & Mantegna, R. N. (2005). A tool for filtering information in complex systems. PNAS, 102(30), 10421-10426.
Examples
>>> import networkx as nx >>> from networkx_backbone import boolean_filter, planar_maximally_filtered_graph >>> G = nx.les_miserables_graph() >>> H = planar_maximally_filtered_graph(G) >>> backbone = boolean_filter(H, "pmfg_keep") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(mn + m log m)time andO(n + m)space.
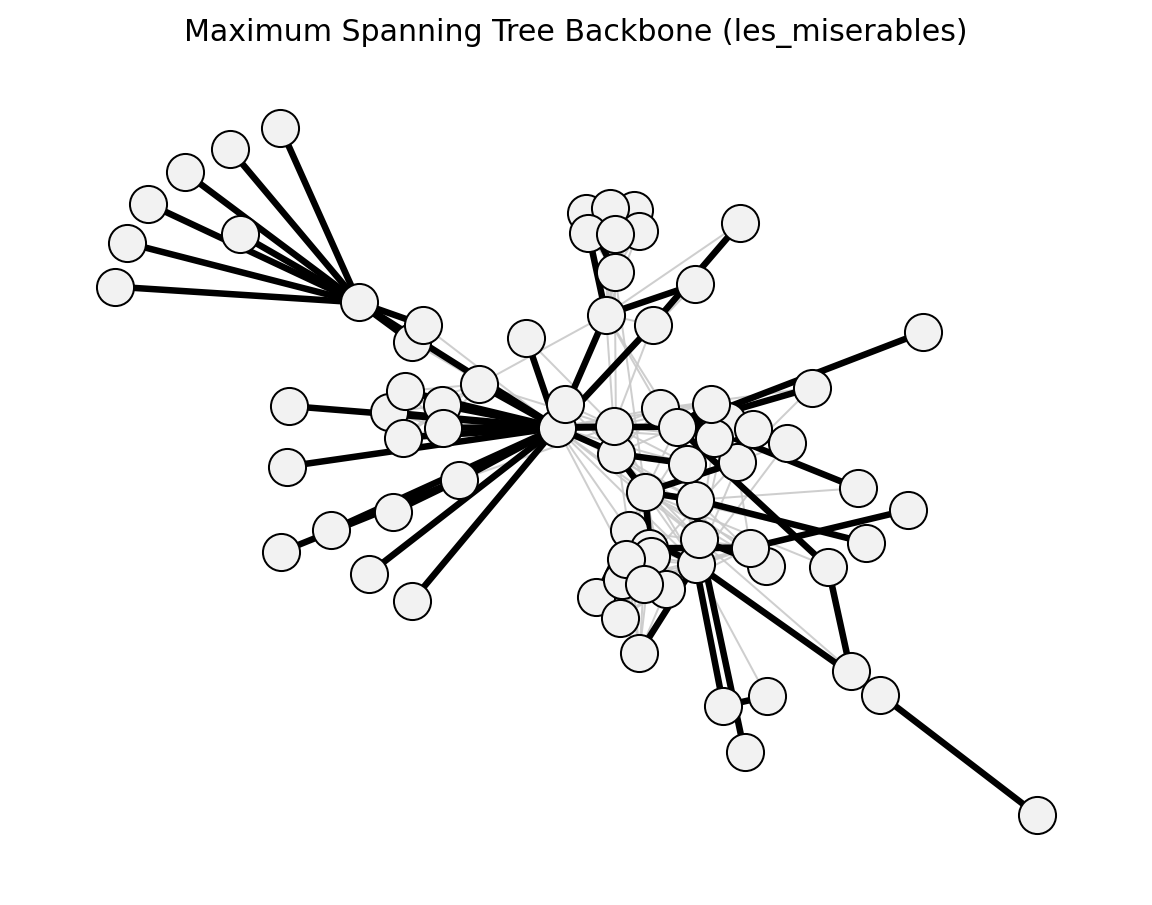
- networkx_backbone.maximum_spanning_tree_backbone(G, weight='weight')[source]#
Extract the maximum spanning tree as a backbone.
- Parameters:
G (graph) – A NetworkX graph.
weight (string, optional (default="weight")) – Edge attribute key used as the weight.
- Returns:
H – A copy of G with boolean
"mst_keep"on edges.- Return type:
graph
Examples
>>> import networkx as nx >>> from networkx_backbone import boolean_filter, maximum_spanning_tree_backbone >>> G = nx.les_miserables_graph() >>> H = maximum_spanning_tree_backbone(G) >>> backbone = boolean_filter(H, "mst_keep") >>> backbone.number_of_edges() == G.number_of_nodes() - 1 True
Computational complexity estimate is
O(m log n)time andO(n + m)space.
Gallery Examples#
Function Image Reference

planar_maximally_filtered_graph()
maximum_spanning_tree_backbone()