Statistical Methods#
Examples in this module use nx.les_miserables_graph().
All functions return scored full graphs; apply
threshold_filter() or
boolean_filter() as the second step.
Complexity classes are provided in each function docstring.
Statistical backbone extraction methods.
These methods evaluate edge significance using hypothesis testing against
a null model. Each method computes a p-value (or score) for every edge,
which can then be filtered using functions in networkx_backbone.filters.
- networkx_backbone.statistical.disparity_filter()[source]#
Serrano et al. (2009) – uniform distribution null model.
- networkx_backbone.statistical.noise_corrected_filter()[source]#
Coscia & Neffke (2017) – Bayesian binomial framework.
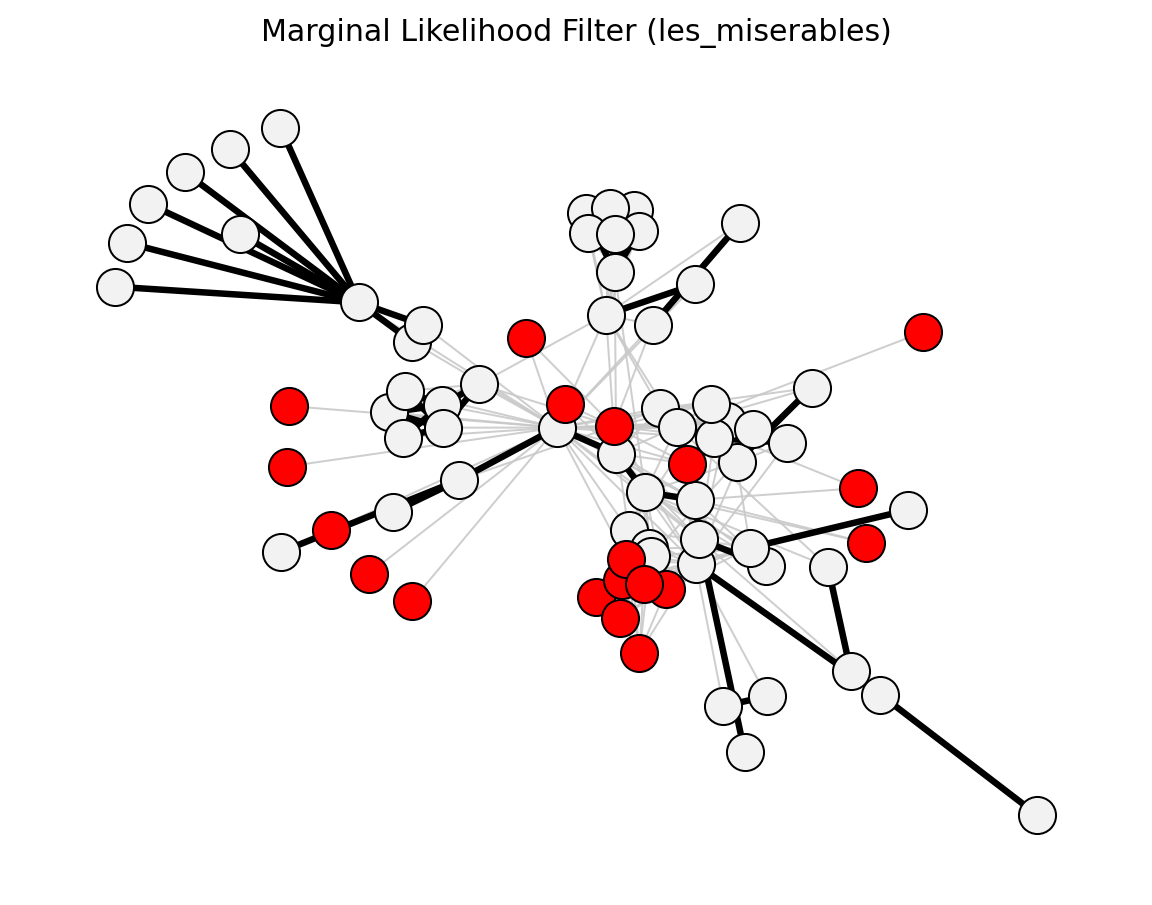
- networkx_backbone.statistical.marginal_likelihood_filter()[source]#
Dianati (2016) – binomial null considering both endpoints.
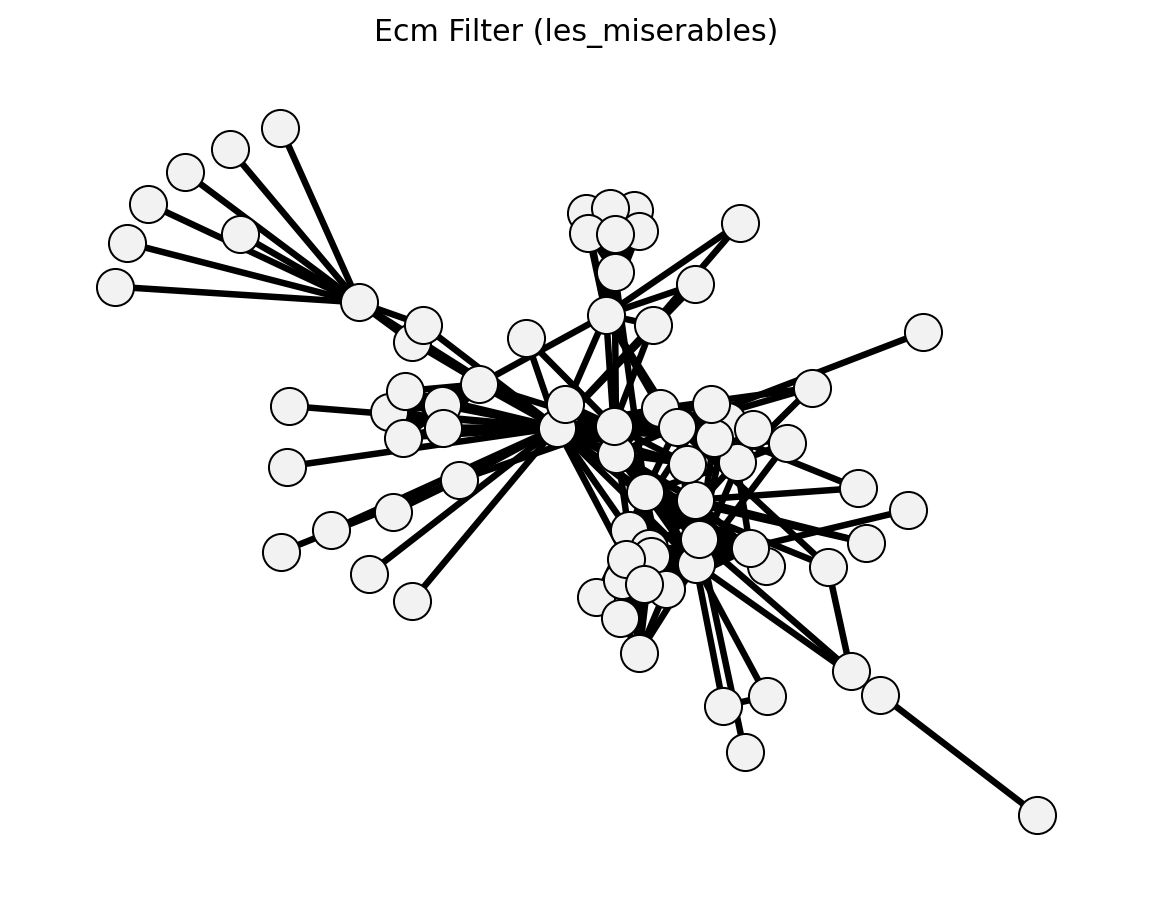
- networkx_backbone.statistical.ecm_filter()[source]#
Gemmetto et al. (2017) – enhanced configuration model.
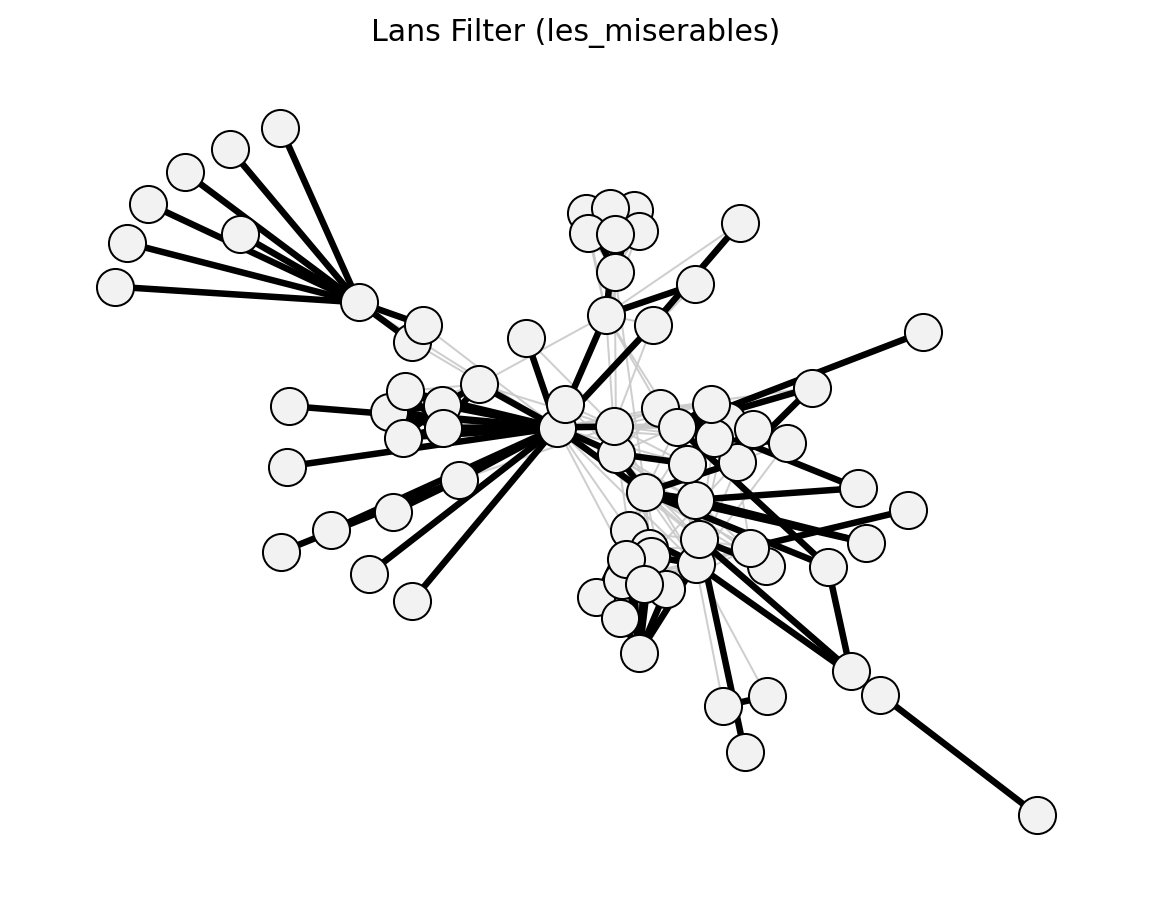
- networkx_backbone.statistical.lans_filter()[source]#
Foti et al. (2011) – nonparametric, empirical CDF-based.
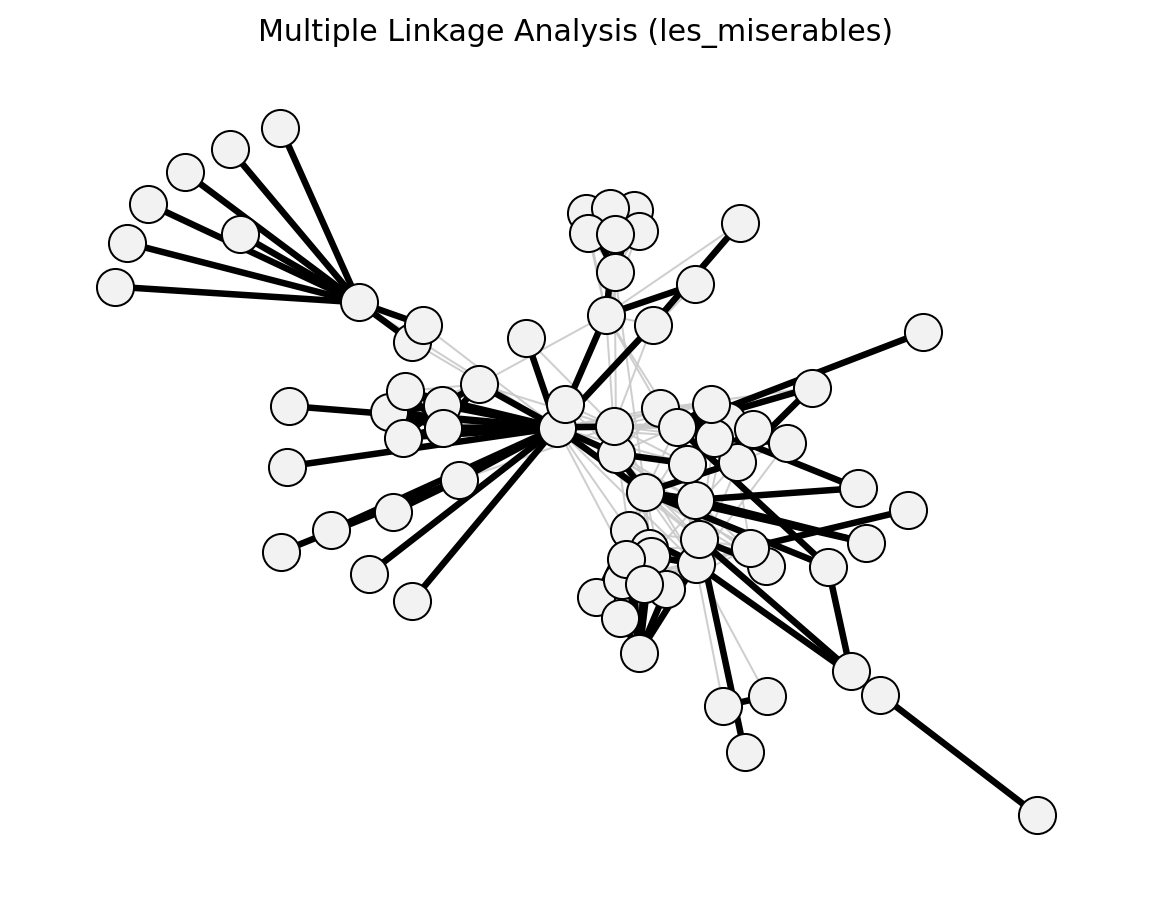
- networkx_backbone.statistical.multiple_linkage_analysis()[source]#
Local linkage significance backbone extraction.
- networkx_backbone.disparity_filter(G, weight='weight')[source]#
Compute disparity filter p-values for each edge.
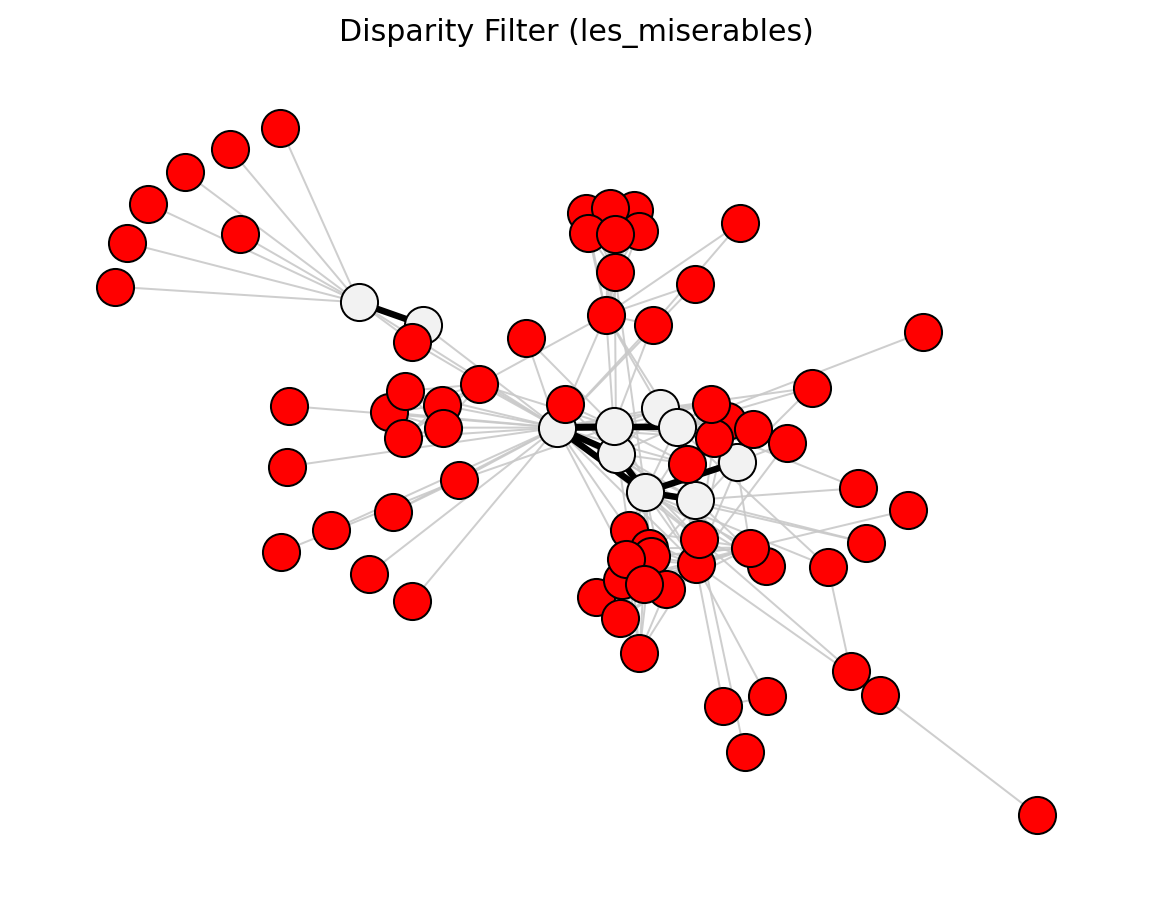
The disparity filter [1] tests whether an edge’s weight is statistically significant compared to a null model where each node’s total strength is uniformly distributed across its edges.
For each node u with degree k_u and strength s_u, the normalised weight of edge (u, v) is
p_uv = w_uv / s_u. Under the null the probability of observing a normalised weight >= p_uv is:alpha_uv = (1.0 - p_uv) ** (k_u - 1)
For an undirected edge the p-value is the minimum of the values computed from each endpoint. For a directed edge the p-value is computed from the source node only (out-strength, out-degree).
- Parameters:
G (networkx.Graph or networkx.DiGraph) – A NetworkX graph.
weight (string, optional (default="weight")) – Edge attribute key for weights. All weights must be positive.
- Returns:
H – A copy of G (same type) with
"disparity_pvalue"added as an edge attribute.- Return type:
graph
- Raises:
NetworkXError – If any edge has a non-positive or missing weight.
References
[1]Serrano, M. A., Boguna, M., & Vespignani, A. (2009). Extracting the multiscale backbone of complex weighted networks. PNAS, 106(16), 6483-6488.
Examples
>>> import networkx as nx >>> from networkx_backbone import disparity_filter, threshold_filter >>> G = nx.les_miserables_graph() >>> H = disparity_filter(G) >>> backbone = threshold_filter(H, "disparity_pvalue", 0.05, mode="below") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(n + m)time andO(n + m)space.
- networkx_backbone.noise_corrected_filter(G, weight='weight')[source]#
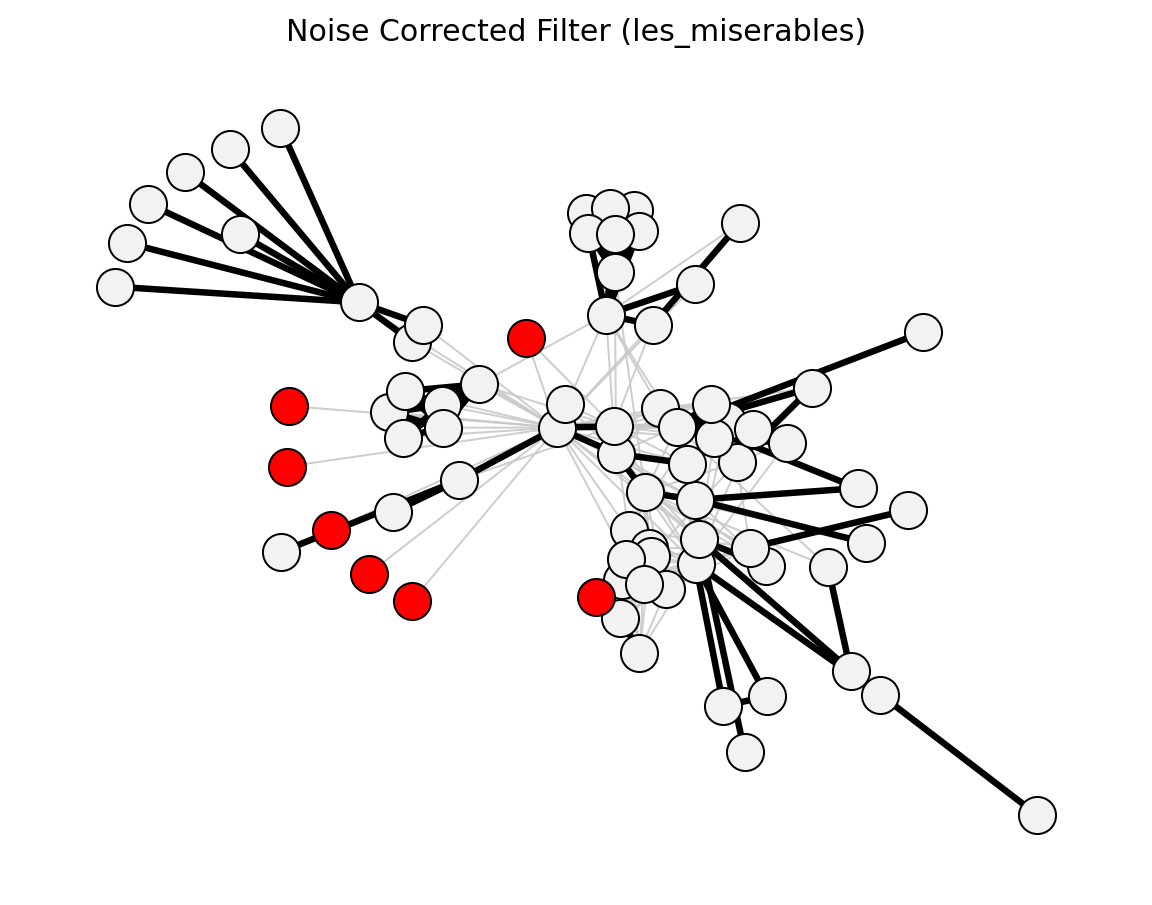
Compute noise-corrected edge significance scores.
Uses a Bayesian framework where each edge weight is modelled as the outcome of a binomial process [1]. The expected weight of edge (u, v) given a total network weight W is:
E[w_uv] = (s_u * s_v) / W
The score measures how many standard deviations the observed weight lies above the expectation (a z-score). Higher scores indicate more significant edges.
For directed graphs, out-strength of u and in-strength of v are used.
- Parameters:
G (networkx.Graph or networkx.DiGraph) – A NetworkX graph.
weight (string, optional (default="weight")) – Edge attribute key for weights. All weights must be positive.
- Returns:
H – A copy of G (same type) with
"nc_score"edge attribute (z-score; higher means more significant).- Return type:
graph
- Raises:
NetworkXError – If any edge has a non-positive or missing weight.
References
[1]Coscia, M. & Neffke, F. M. (2017). Network backboning with noisy data. Proc. IEEE ICDE, 425-436.
Examples
>>> import networkx as nx >>> from networkx_backbone import noise_corrected_filter, threshold_filter >>> G = nx.les_miserables_graph() >>> H = noise_corrected_filter(G) >>> backbone = threshold_filter(H, "nc_score", 2.0, mode="above") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(n + m)time andO(n + m)space.
- networkx_backbone.marginal_likelihood_filter(G, weight='weight')[source]#
Compute marginal likelihood p-values for each edge.
The marginal likelihood filter [1] considers edge weights as integer counts under a binomial null model that accounts for both endpoints’ strengths.
- Parameters:
G (networkx.Graph or networkx.DiGraph) – A NetworkX graph. Integer weights are recommended.
weight (string, optional (default="weight")) – Edge attribute key for weights. All weights must be positive.
- Returns:
H – A copy of G (same type) with
"ml_pvalue"edge attribute.- Return type:
graph
- Raises:
NetworkXError – If any edge has a non-positive or missing weight.
References
[1]Dianati, N. (2016). Unwinding the hairball graph: Pruning algorithms for weighted complex networks. Physical Review E, 93, 012304.
Examples
>>> import networkx as nx >>> from networkx_backbone import marginal_likelihood_filter, threshold_filter >>> G = nx.les_miserables_graph() >>> H = marginal_likelihood_filter(G) >>> backbone = threshold_filter(H, "ml_pvalue", 0.05, mode="below") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(n + m)time andO(n + m)space.
- networkx_backbone.ecm_filter(G, weight='weight', max_iter=1000, tol=1e-06)[source]#
Compute ECM (Enhanced Configuration Model) p-values.
The ECM [1] is a maximum-entropy null model for weighted networks that preserves the expected degree and strength sequence. Lagrange multipliers are assigned to each node and solved iteratively.
- Parameters:
G (networkx.Graph or networkx.DiGraph) – A NetworkX graph.
weight (string, optional (default="weight")) – Edge attribute key for weights. All weights must be positive.
max_iter (int, optional (default=1000)) – Maximum number of iterations for the fixed-point solver.
tol (float, optional (default=1e-6)) – Convergence tolerance for the Lagrange multipliers.
- Returns:
H – A copy of G (same type) with
"ecm_pvalue"edge attribute.- Return type:
graph
- Raises:
NetworkXError – If any edge has a non-positive or missing weight.
References
[1]Gemmetto, V., Cardillo, A., & Garlaschelli, D. (2017). Irreducible network backbones: unbiased graph filtering via maximum entropy. arXiv:1706.00230.
Examples
>>> import networkx as nx >>> from networkx_backbone import ecm_filter, threshold_filter >>> G = nx.les_miserables_graph() >>> H = ecm_filter(G) >>> backbone = threshold_filter(H, "ecm_pvalue", 0.05, mode="below") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(I * n^2 + m)time andO(n + m)space.
- networkx_backbone.lans_filter(G, weight='weight')[source]#
Compute LANS (Locally Adaptive Network Sparsification) p-values.
LANS [1] is a nonparametric method that makes no distributional assumptions. For each edge (u, v), it computes the fraction of edges at node u (and node v) that have weight <= w_uv — the empirical CDF value. The edge p-value is one minus the maximum of the two eCDF values (so lower p means more significant).
For directed graphs, only the source node’s out-edges are used.
- Parameters:
G (networkx.Graph or networkx.DiGraph) – A NetworkX graph.
weight (string, optional (default="weight")) – Edge attribute key for weights. All weights must be positive.
- Returns:
H – A copy of G (same type) with
"lans_pvalue"edge attribute.- Return type:
graph
- Raises:
NetworkXError – If any edge has a non-positive or missing weight.
References
[1]Foti, N. J., Hughes, J. M., & Rockmore, D. N. (2011). Nonparametric sparsification of complex multiscale networks. PLoS ONE, 6(2), e16431.
Examples
>>> import networkx as nx >>> from networkx_backbone import lans_filter, threshold_filter >>> G = nx.les_miserables_graph() >>> H = lans_filter(G) >>> backbone = threshold_filter(H, "lans_pvalue", 0.05, mode="below") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(m log n)time andO(n + m)space.
- networkx_backbone.multiple_linkage_analysis(G, alpha=0.05, weight='weight')[source]#
Extract a backbone using Multiple Linkage Analysis (MLA).
MLA is a local-significance method that selects edges whose weights are unusually high relative to neighboring edges. This implementation uses edge-level empirical CDF p-values from
lans_filter()and retains edges with p-value <=alpha.- Parameters:
G (networkx.Graph or networkx.DiGraph) – A NetworkX graph.
alpha (float, optional (default=0.05)) – Significance threshold in [0, 1].
weight (string, optional (default="weight")) – Edge attribute key for weights. All weights must be positive.
- Returns:
H – A copy of G with
"lans_pvalue","mla_pvalue", and boolean"mla_keep"edge attributes.- Return type:
graph
- Raises:
ValueError – If
alphais outside [0, 1].NetworkXError – If any edge has a non-positive or missing weight.
References
[1]Van Nuffel, N., Heyndrickx, C., & Wets, G. (2010). Measuring hierarchy and reciprocity in networks.
[2]Yassin, A., Haidar, A., Cherifi, H., Seba, H., & Togni, O. (2023). An evaluation tool for backbone extraction techniques in weighted complex networks. Scientific Reports, 13, 17000.
Examples
>>> import networkx as nx >>> from networkx_backbone import boolean_filter, multiple_linkage_analysis >>> G = nx.les_miserables_graph() >>> H = multiple_linkage_analysis(G, alpha=0.5) >>> backbone = boolean_filter(H, "mla_keep") >>> backbone.number_of_edges() <= H.number_of_edges() True
Computational complexity estimate is
O(m log n)time andO(n + m)space.
Gallery Examples#
Function Image Reference
Alias Names
- networkx_backbone.disparity(G, weight='weight')[source]#
Alias for
disparity_filter().Computational complexity estimate is
O(n + m)time andO(n + m)space.
- networkx_backbone.mlf(G, weight='weight')[source]#
Alias for
marginal_likelihood_filter().Computational complexity estimate is
O(n + m)time andO(n + m)space.
- networkx_backbone.lans(G, weight='weight')[source]#
Alias for
lans_filter().Computational complexity estimate is
O(m log n)time andO(n + m)space.